RiboCop rRNA Depletion Kit for Yeast
Why limit yourself to mRNA, when you can analyze also the non-coding transcriptome? Unleash the Yeast!
RiboCop for Yeast
Yeast total RNA is comprised of large amounts of undesired ribosomal RNA (rRNA) which accounts for up to ~97 % of all transcripts. Lexogen’s RiboCop rRNA Depletion Kits for Yeast remove undesired cytoplasmic (25S, 18S, 5.8S, 5S, and 35S) and mitochondrial (21S, 15S, ACI60_gr01 and ACI60_gr02) rRNA from intact as well as degraded input RNA. Ribo-depletion affords an unbiased view of the transcriptome, including non-coding and non-polyadenylated RNA species.
RiboCop for Yeast is ideally suited for Next Generation Sequencing (NGS) approaches in combination with CORALL RNA-Seq V2 Library Preparation Kits. RiboCop Kits are also compatible with other random primed total RNA library prep kits.
Performance
Robust Performance Over a Wide Range of Input Amounts
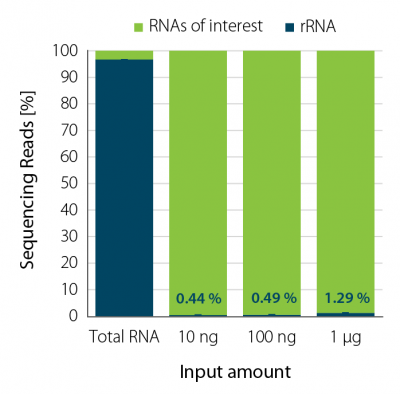
Ribosomal RNA was depleted from Saccharomyces cerevisiae total RNA using RiboCop for Yeast over a wide range of input amounts (10 ng to 1 µg). RiboCop for Yeast efficiently reduces rRNA reads to ~1 % for all input amounts (Fig. 1).
Figure 1 | RiboCop rRNA Depletion for Yeast efficiently removes rRNA across a wide range of input amounts. NGS libraries were prepared using Lexogen’s CORALL RNA-Seq V2 Library Prep Kit. Successful depletion was monitored by sequencing (NextSeq500, 1×50 bp) and subsequent analysis of remaining rRNA reads from untreated (Total RNA) and depleted S. cerevisiae RNA (10 ng, 100 ng, and 1 µg). The percentage of reads mapping to rRNA is plotted in blue.
RiboCop for Yeast Maintains Unbiased Expression Profiles
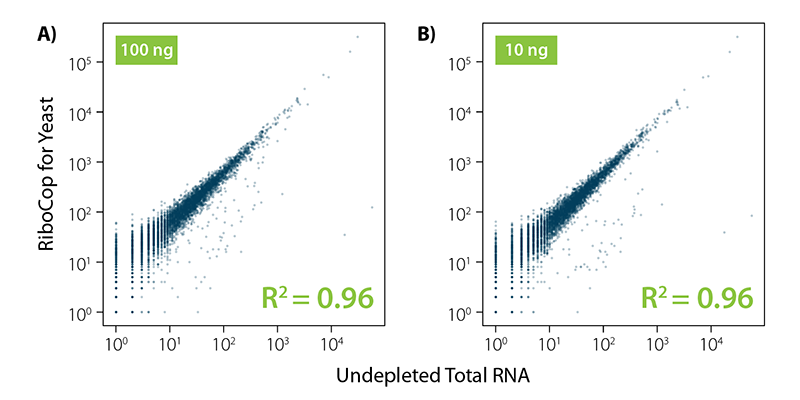
RiboCop for Yeast efficiently removes rRNA while maintaining unbiased transcript expression profiles as indicated by high correlation between transcript abundance in untreated vs. ribo-depleted samples for two different input amounts (Fig. 2).
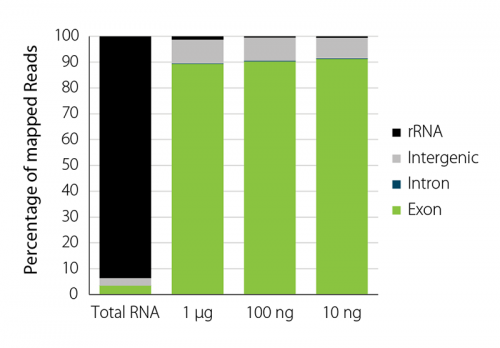
Sequencing reads are focused on RNAs of interest significantly increasing exonic reads for depleted yeast RNA regardless of the input RNA amount (Fig. 3).
Figure 2 | RiboCop for Yeast maintains unbiased expression profiles while efficiently removing undesired ribosomal RNA. Correlation of transcript abundance in untreated samples vs. samples depleted with RiboCop for Yeast using 10 ng and 100 ng S. cerevisiae RNA input. Libraries were prepared using Lexogen’s CORALL RNA-Seq V2 Library Prep Kit and sequenced on Illumina NextSeq (1×50 bp). Reads were mapped against the S.cerevisiae reference genome using STAR aligner and counted with FeatureCounts. Correlations are shown for uniquely mapping reads.
Figure 3 | Feature distribution of mapped reads for S. cerevisiae with and without ribo-depletion. RiboCop for Yeast rRNA depletion significantly increases the fraction of reads mapped to exons. Yeast RNA was processed and sequenced as described in Fig. 2.
Increased Gene Detection upon rRNA Depletion
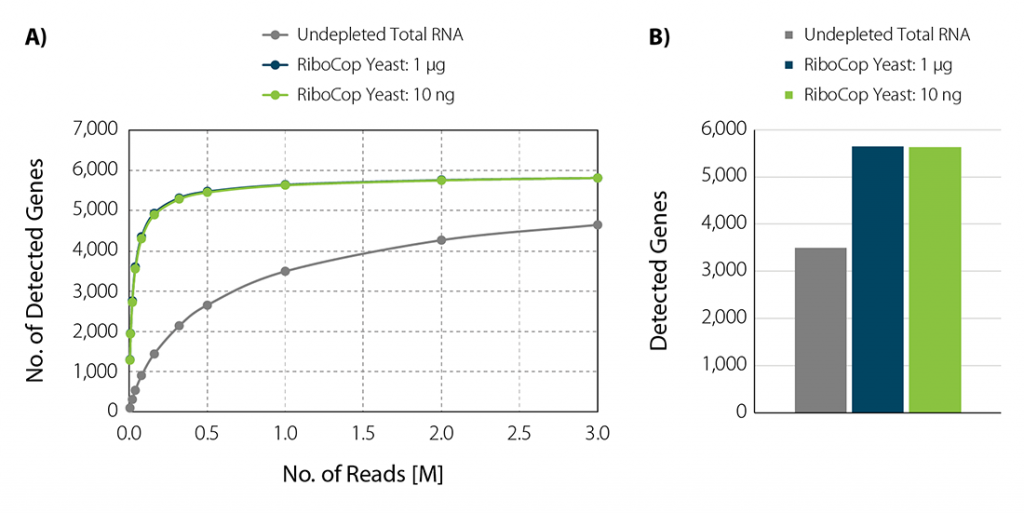
Sequence what matters to your experiment. RiboCop for Yeast affords an unobstructed view of the yeast transcriptome and increases gene detection through efficient rRNA removal (Fig. 4).
Figure 4 | Increased gene detection upon rRNA depletion. A) The number of detected genes per number of reads uniquely mapping to exons (FeatureCounts) was plotted for undepleted total RNA as well as depleted S. cerevisiae RNA across different input amounts. B) Bar plot illustrating the number of detected genes at 1 M reads /sample. S. cerevisiae samples depleted with RiboCop for Yeast using either 1 µg or 10 ng input RNA. Libraries were prepared using Lexogen’s CORALL RNA-Seq V2 Library Prep Kit and sequenced on Illumina NextSeq (1×50 bp). Reads were mapped against the S. cerevisiae reference genome using STAR aligner and counted with FeatureCounts.
Excellent Reproducibility Between Replicates
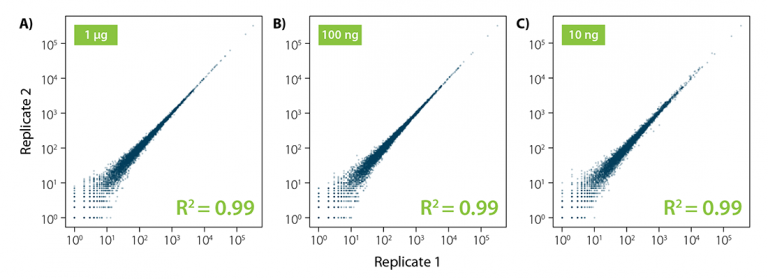
RiboCop protocols are robust and highly reproducible. Correlation plots show excellent reproducibility between replicates independent of the input amount used for the depletion reaction (Fig. 5).
Figure 5 | Excellent reproducibility between replicates. Correlation of replicates for two independent depletion reactions using 10 ng, 100 ng, and 1 µg S. cerevisiae total RNA as input. Total RNA was ribo-depleted with RiboCop for Yeast. Libraries were prepared using Lexogen’s CORALL RNA-Seq V2 Library Prep Kit and sequenced on Illumina NextSeq (1×50 bp). Reads were mapped against the S. cerevisiae reference genome using STAR aligner and counted with FeatureCounts. Correlations are shown including multimapping reads.
RiboCop for Yeast Maintains Undisturbed Expression Profiles Across Various Inputs
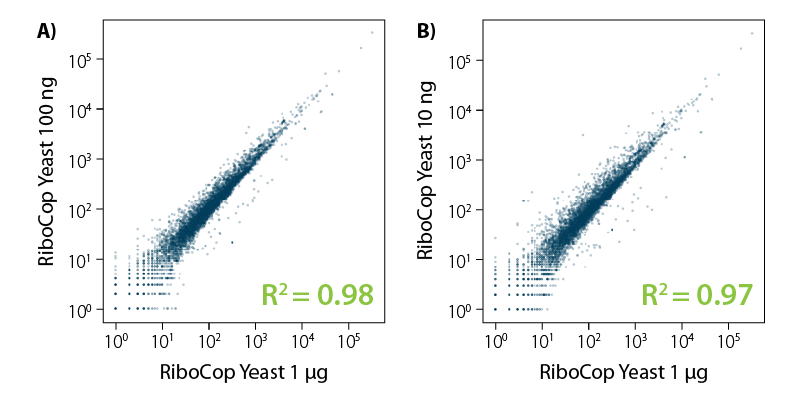
RiboCop protocols are highly reproducible across various input amounts. Correlation plots show consistent transcript profiles and high correlation over a broad range of inputs (Fig. 6).
Figure 6 | RiboCop for Yeast maintains constant transcript abundance across various input amounts. Correlation of transcript abundance for S. cerevisiae samples depleted with RiboCop for Yeast using either 1 µg, 100 ng RNA or 10 ng RNA. Libraries were prepared using Lexogen’s CORALL RNA-Seq V2 Library Prep Kit and sequenced on Illumina NextSeq (1×50 bp). Reads were mapped against the S. cerevisiae reference genome using STAR aligner and counted with FeatureCounts. Correlations are shown including multimapping reads.
Workflow
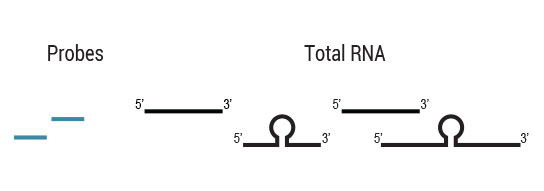
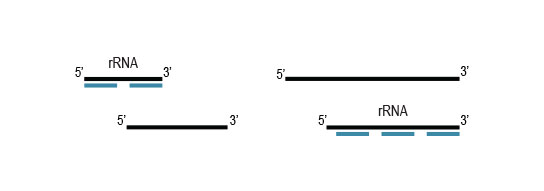
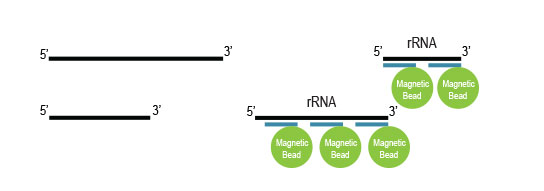
Samples are treated using a set of affinity probes for specific depletion of rRNA sequences. RiboCop probes efficiently remove rRNA and therefore afford a comprehensive view of transcriptome composition.
1hr 30min
30 min
FAQ
Frequently Asked Questions
Access our frequently asked question (FAQ) resources via the buttons below.
Please also check our General Guidelines and FAQ resources!
How do you like the new online FAQ resource? Please share your feedback with us!
Downloads
Safety Data Sheet
If you need more information about our products, please contact us through support@lexogen.com or directly under +43 1 345 1212-41.
Ordering information
| Cat. No. | Product Name |
| 190.24 | RiboCop rRNA Depletion Kit for Yeast, 24 preps |
| 190.96 | RiboCop rRNA Depletion Kit for Yeast, 96 preps |
First time user of RiboCop?
First Time User? We’re excited to offer you an exclusive introductory offer.
Buy from our Webstore
Need a web quote?
You can generate a web quote by Register or Login to your account. In the account settings please fill in your billing and shipping address. Add products to your cart, view cart and click the “Generate Quote” button. A quote in PDF format will be generated and ready to download. You can use this PDF document to place an order by sending it directly to sales@lexogen.com.
Web quoting is not available for countries served by our distributors. Please contact your local distributor for a quote.