SPLIT RNA Extraction Kit -
Purify RNA to the highest grade!
SPLIT RNA Extraction Kit
The SPLIT RNA Extraction Kit enables fast and highly efficient extraction of high-quality, high-purity RNA from various biological samples, including cell culture, animal and plant tissue, and fluid samples. The obtained RNA is ideal for seamless library preparation for Next Generation Sequencing and other demanding applications such as full-length reverse transcription, sample preparation for microarray analysis, or RT-qPCR. SPLIT recovers the complete RNA size range, including small RNAs (<200 nt). Additionally, large and small RNA-enriched fractions can be extracted by following a supplemental protocol.
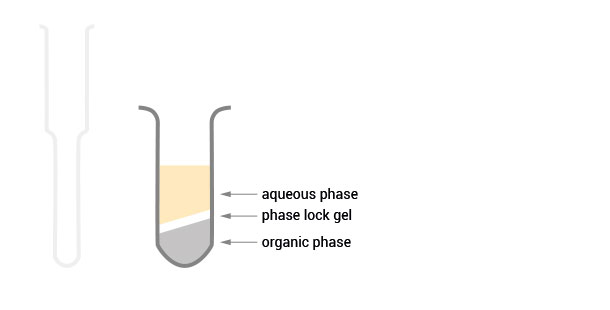
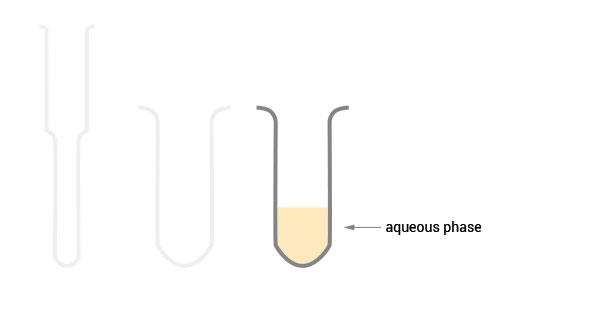
SPLIT RNA Extraction Kit (Cat. No. 008) offers universal, species-independent RNA extraction from a broad range of materials, including animal and plant tissue, cell cultures, liquid samples, and FFPE samples. SPLIT’s streamlined protocol delivers high-purity RNA within 30 minutes using Phase Lock Gel tubes for rapid, convenient, and safe phase separation.
Performance
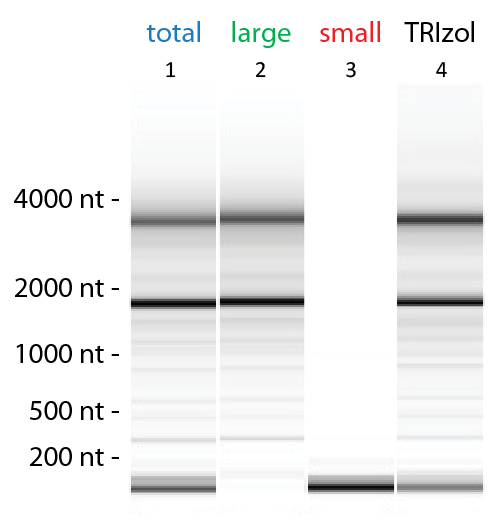
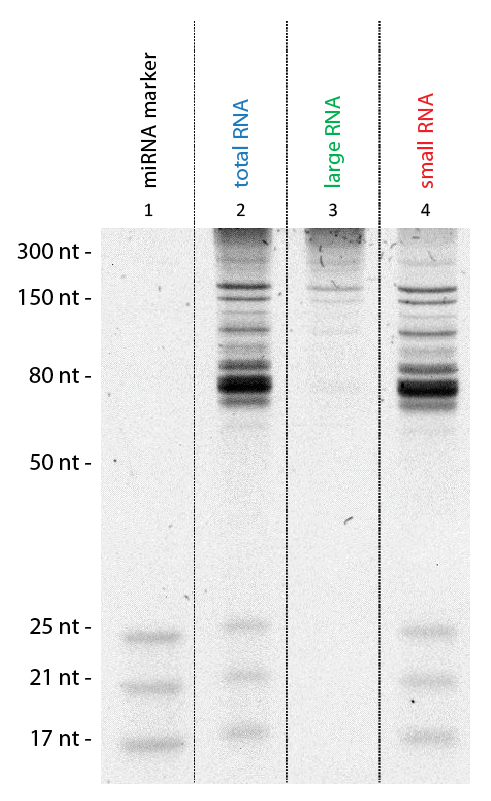
Exceptional Yields and High-quality RNA Extraction
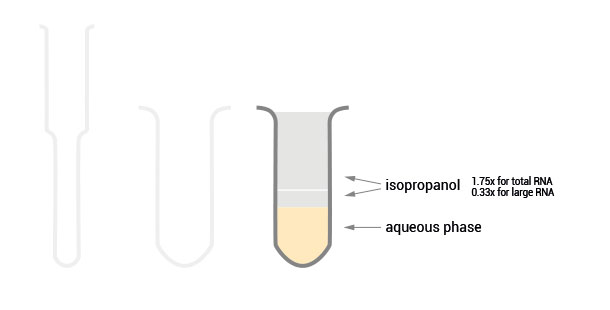
Flexible RNA Isolation of Total RNA or Size-enriched RNA Fractions
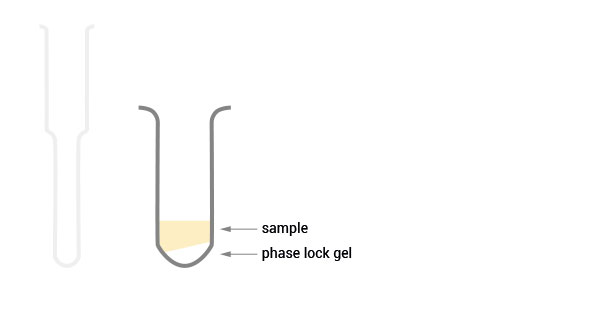
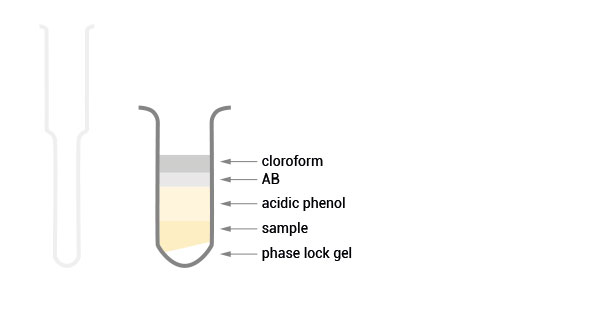
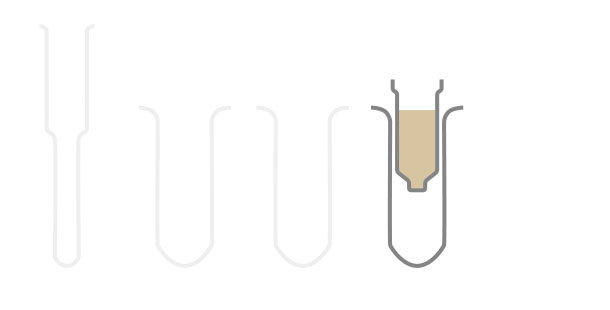
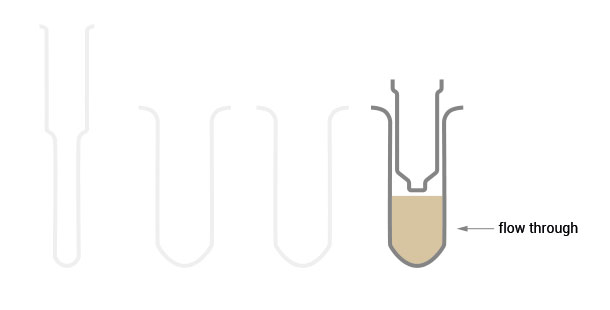
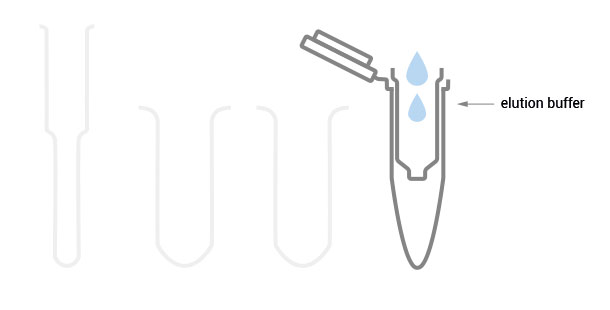
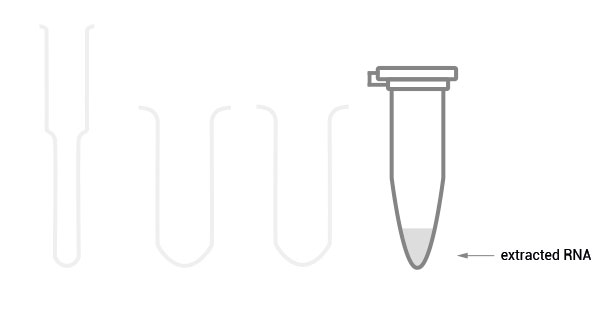
Workflow
30 min
FAQ
Frequently Asked Questions
Access our frequently asked question (FAQ) resources via the buttons below.
Please also check our General Guidelines and FAQ resources!
How do you like the new online FAQ resource? Please share your feedback with us!
Troubleshooting Guide
View the Troubleshooting Guide
| Problem | Likely cause | Comments and suggestions |
| No or poor phase separation | Cooled phase-lock gel or insufficient centrifugation |
|
| Incorrect cut-off | Volume of isopropanol addition |
|
| Degraded RNA | RNA source |
|
| RNase contamination |
| |
| Low absorption ratios (i.e., peak at 230 nm) | Contamination with organic solvents, salts, or metal ions |
|
| Low or no RNA yield | RNA remains on spin column |
|
| Spin column was overloaded |
| |
| Problem in downstream application | Salt or ethanol carry-over during elution |
|
Downloads
Explore our dedicated FFPE RNA extraction kit: SPLIT One-step FFPE RNA Extraction Kit
Safety Data Sheet
If you need more information about our products, please contact us through support@lexogen.com or directly under +43 1 345 1212-41.
Ordering Information
| Cat. No. | Product Name |
| 008.48 | SPLIT RNA Extraction Kit, 48 extractions |
First time user of SPLIT?
First Time User? We’re excited to offer you an exclusive introductory offer.
Buy from our Webstore
Need a web quote?
You can generate a web quote by Register or Login to your account. In the account settings please fill in your billing and shipping address. Add products to your cart, view cart and click the “Generate Quote” button. A quote in PDF format will be generated and ready to download. You can use this PDF document to place an order by sending it directly to sales@lexogen.com.
Web quoting is not available for countries served by our distributors. Please contact your local distributor for a quote.