Lexogen 12 nt UDI Sets -
Tag your libraries with ultimate accuracy
Lexogen 12 nt Unique Dual Index (UDI) System for RNA-Seq
Lexogen’s 12 nt Unique Dual Indices (UDIs) are introduced at the PCR step and enable unique barcoding of up to 384 samples per lane run. Lexogen’s 12 nt UDIs feature superior error correction for maximal sequencing data output.
Lexogen’s 12 nt UDIs are available as convenient bundles or Add-on sets, compatible with the following Lexogen library prep solutions:
Performance
Ensuring Accurate RNA-Seq Data with Unique Dual Indexes
Superior Error Correction Maximizes Sequencing Yield
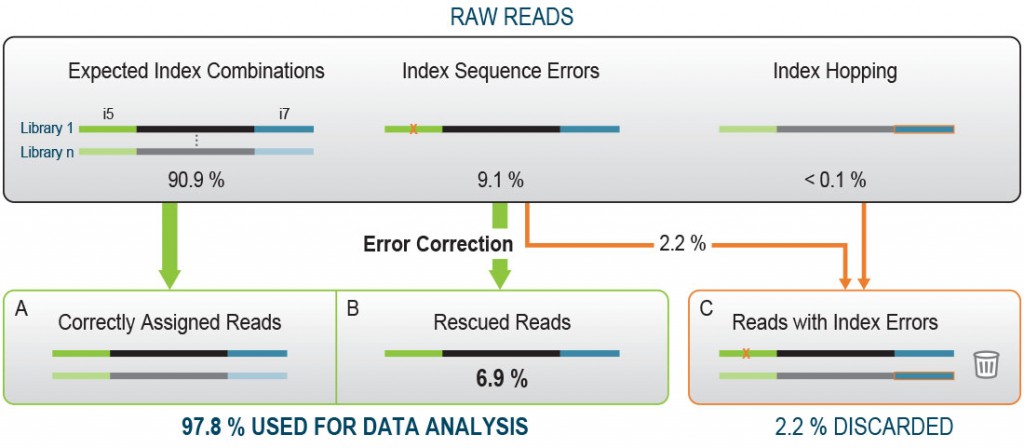
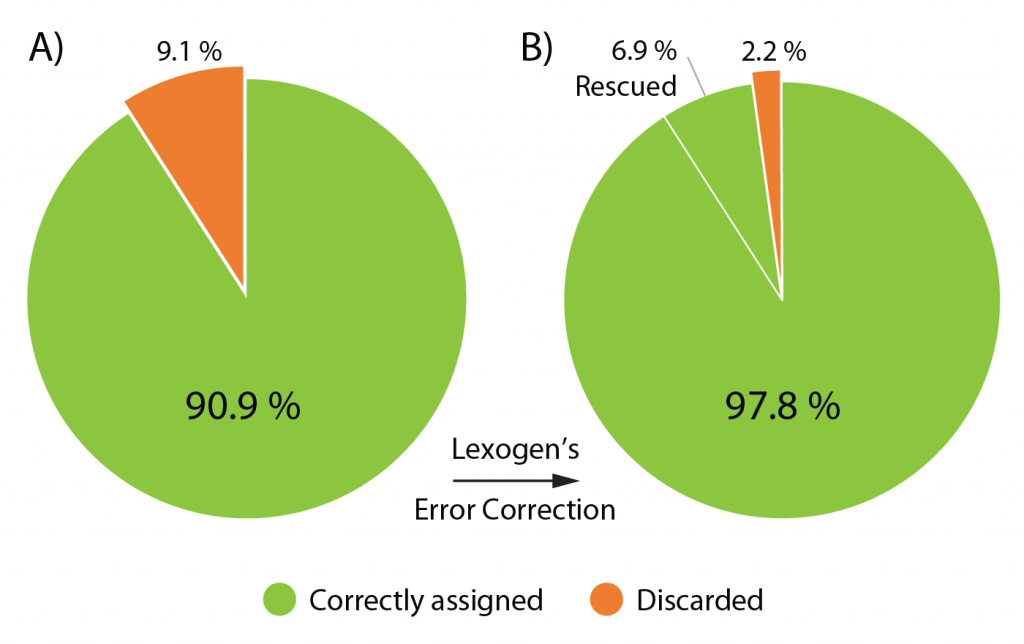
The Lexogen UDI 12 nt Unique Dual Indices are 12 nucleotides (nt) long and designed to maximize inter-index distance for different sample numbers and index read-out lengths. In a typical experiment using the full 12 nt index read-out, around 9.1 % of the initial raw reads contain a random Index Sequence Error (Fig. 2A). This renders them undetermined, hence removing these reads from downstream analysis.
Lexogen’s advanced index design enables the rescue of 76 % of these undetermined reads increasing the useful output thereby increases to 97.8 % of the initial reads, an unprecedented performance due to the cutting-edge index design (Fig. 2B).
Scalable Index Read-out Length
Lexogen’s 12 nt Unique Dual Indexes (UDIs) offer flexible read-out lengths of 8, 10, or 12 nucleotides for unique barcoding and to accommodate diverse multiplexing needs, ensuring accuracy and reliable error correction based on experiment type, sequencing equipment, desired read depth, and / or the number of pooled libraries. Longer 12 nt UDIs, while requiring slightly more sequencing cycles, provide enhanced error correction capabilities for more complex index sequence errors, eliminating the need for multiple indexing sets.
Nested Index Set Design for Highest Accuracy
Highest Inter-index Distance for Maximal Error Correction Capacity
| Subset | 12 nt | 10 nt | 8 nt | |||
| D | ec | D | ec | D | ec | |
| 384 | 4 | 1 | 3 | 1 | 2 | 1* |
| └ 96 | 5 | 2 | 4 | 1 | 3 | 1 |
| └ 24 | 6 | 2 | 5 | 2 | 4 | 1 |
| └ 4 | 7 | 3 | 6 | 2 | 5 | 2 |
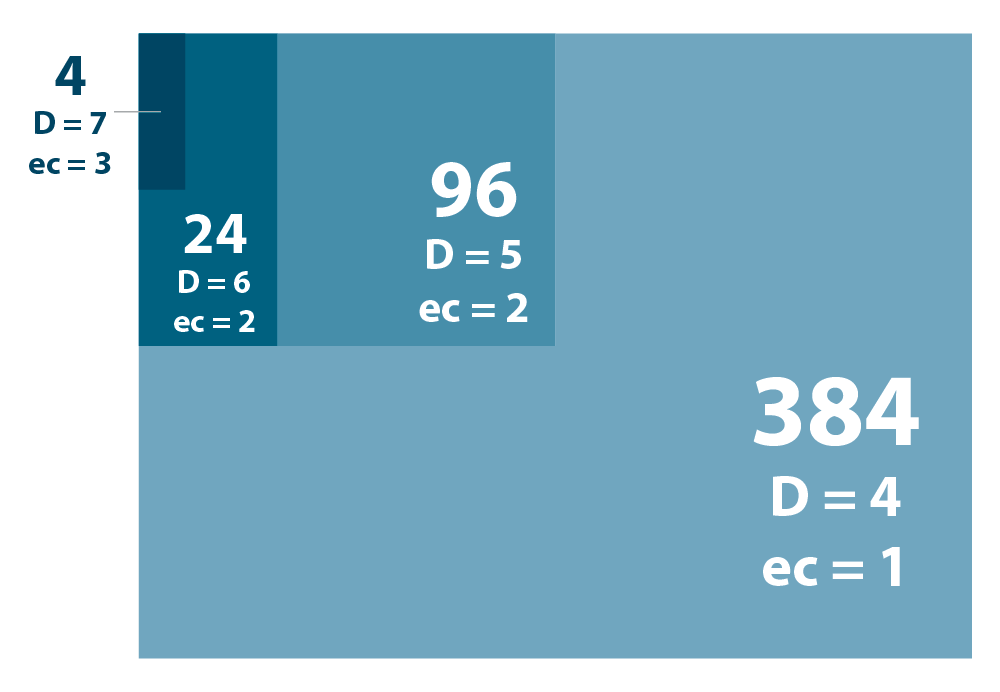
Table 1 | Comparison of distance and error correction capacity in Lexogen’s nested 12 nt UDI sets with 8, 10, and 12 nt read-out. Inter-index distance (D) and number of errors that can be corrected (ec) are compared for subsets of 384, 96, 24, and 4 libraries and the three possible read-out lengths. For smaller subsets (up to 96 samples) a read-out of 8 or 10 nt allows correction of one error and thus recovery of additional reads. Larger subsets require a read-out of 10 or 12 nt to benefit from the error correction. The ec values represent the number of all errors (including substitutions, insertions, and deletions) that can be confidently corrected, except for *. In this case error correction can only address substitutions.
FAQ
Frequently Asked Questions
Access our frequently asked question (FAQ) resources via the buttons below.
Please also check our General Guidelines and FAQ resources!
How do you like the new online FAQ resource? Please share your feedback with us!
Downloads
Lexogen 12 nt Unique Dual Index System (UDI) for RNA-Seq
New Version (Version 2):
Older Version (Version 1):
Safety Data Sheet
If you need more information about our products, please contact us through support@lexogen.com or directly under +43 1 345 1212-41.
Ordering Information
QuantSeq FWD - Catalog Numbers
| Cat. No. | Product Name |
| QuantSeq FWD | |
| 191.96 | QuantSeq 3’ mRNA-Seq V2 Library Prep Kit FWD with UDI 12 nt Set A1, (UDI12A_0001-0096), 1 rxn/UDI |
| 192.24 | QuantSeq 3’ mRNA-Seq V2 Library Prep Kit FWD with UDI 12 nt Set B1, (UDI12B_0001-0024), 1 rxn/UDI |
| 192.96 | QuantSeq 3’ mRNA-Seq V2 Library Prep Kit FWD with UDI 12 nt Set B1, (UDI12B_0001-0096), 1 rxn/UDI |
| 193.384 | QuantSeq 3’ mRNA-Seq V2 Library Prep Kit FWD with UDI 12 nt Sets A1-A4, (UDI12A_0001-0384), 1 rxn/UDI |
| 194.96 | QuantSeq 3’ mRNA-Seq V2 Library Prep Kit FWD with UDI 12 nt Set A2, (UDI12A_0097-0192), 1 rxn/UDI |
| 195.96 | QuantSeq 3’ mRNA-Seq V2 Library Prep Kit FWD with UDI 12 nt Set A3, (UDI12A_0193-0288), 1 rxn/UDI |
| 196.96 | QuantSeq 3’ mRNA-Seq V2 Library Prep Kit FWD with UDI 12 nt Set A4, (UDI12A_0289-0384), 1 rxn/UDI |
CORALL mRNA-Seq - Catalog Numbers
| Cat. No. | Product Name |
| CORALL mRNA-Seq | |
| 177.96 | CORALL mRNA-Seq V2 Library Prep Kit with UDI 12 nt Set A1, (UDI12A_0001-0096), 96 preps |
| 178.96 | CORALL mRNA-Seq V2 Library Prep Kit with UDI 12 nt Set A2, (UDI12A_0097-0192), 96 preps |
| 179.96 | CORALL mRNA-Seq V2 Library Prep Kit with UDI 12 nt Set A3, (UDI12A_0193-0288), 96 preps |
| 180.96 | CORALL mRNA-Seq V2 Library Prep Kit with UDI 12 nt Set A4, (UDI12A_0289-0384), 96 preps |
| 181.96 | CORALL mRNA-Seq V2 Library Prep Kit with UDI 12 nt Set B1, (UDI12B_0001-0096), 96 preps |
| 182.384 | CORALL mRNA-Seq V2 Library Prep Kit with UDI 12 nt Sets A1-A4 (UDI12A_0001-0384), 384 preps |
CORALL RNA-Seq without rRNA Depletion - Catalog Numbers
| Cat. No. | Product Name |
| CORALL RNA-Seq Stand-alone Kits without rRNA Depletion, including Lexogen 12 nt UDI Sets | |
| 171.24 | CORALL RNA-Seq V2 Library Prep Kit with UDI 12 nt Set A1, (UDI12A_0001-0024), 24 preps |
| 171.96 | CORALL RNA-Seq V2 Library Prep Kit with UDI 12 nt Set A1, (UDI12A_0001-0096), 96 preps |
| 172.96 | CORALL RNA-Seq V2 Library Prep Kit with UDI 12 nt Set A2, (UDI12A_0097-0192), 96 preps |
| 173.96 | CORALL RNA-Seq V2 Library Prep Kit with UDI 12 nt Set A3, (UDI12A_0193-0288), 96 preps |
| 174.96 | CORALL RNA-Seq V2 Library Prep Kit with UDI 12 nt Set A4, (UDI12A_0289-0384), 96 preps |
| 175.24 | CORALL RNA-Seq V2 Library Prep Kit with UDI 12 nt Set B1, (UDI12B_0001-0024), 24 preps |
| 175.96 | CORALL RNA-Seq V2 Library Prep Kit with UDI 12 nt Set B1, (UDI12B_0001-0096), 96 preps |
| 176.384 | CORALL RNA-Seq V2 Library Prep Kit with UDI 12 nt Sets A1-A4 (UDI12A_0001-0384), 384 preps |
CORALL RNA-Seq with rRNA Depletion - Catalog Numbers
| Cat. No. | Product Name |
| CORALL RNA-Seq with rRNA Depletion, including Lexogen 12 nt UDI Sets | 183.24 | RiboCop (HMR) and CORALL Total RNA-Seq V2 Library Prep Kit with UDI 12 nt Set A1, (UDI12A_0001-0024), 24 preps |
| 183.96 | RiboCop (HMR) and CORALL Total RNA-Seq V2 Library Prep Kit with UDI 12 nt Set A1, (UDI12A_0001-0096), 96 preps |
| 184.24 | RiboCop (HMR) and CORALL Total RNA-Seq V2 Library Prep Kit with UDI 12 nt Set B1, (UDI12B_0001-0024), 24 preps |
| 184.96 | RiboCop (HMR) and CORALL Total RNA-Seq V2 Library Prep Kit with UDI 12 nt Set B1, (UDI12B_0001-0096), 96 preps |
LUTHOR HD - Catalog Numbers
| Cat. No. | Product Name |
| LUTHOR HD | |
| 221.24 | LUTHOR High-Definition Single-Cell 3’ mRNA-Seq Kit with UDI 12 nt Set B1, (UDI12B_0001-0024), 24 preps |
| 221.96 | LUTHOR High-Definition Single-Cell 3’ mRNA-Seq Kit with UDI 12 nt Set B1, (UDI12B_0001-0096), 96 preps |
UDI Add-on Kits (UDI plates only), e.g., for QuantSeq-Pool and LUTHOR HD Pool - Catalog Numbers
| Cat. No. | Product Name |
| UDI Add-on Kits (UDI plates only), e.g., for QuantSeq-Pool and LUTHOR HD Pool | |
| 101.96 | Lexogen UDI 12 nt Set A1 (UDI12A_0001-0096), 1 rxn/UDI |
| 102.96 | Lexogen UDI 12 nt Set A2 (UDI12A_0097-0192), 1 rxn/UDI |
| 103.96 | Lexogen UDI 12 nt Set A3 (UDI12A_0193-0288), 1 rxn/UDI |
| 104.96 | Lexogen UDI 12 nt Set A4 (UDI12A_0289-0384), 1 rxn/UDI |
| 105.96 | Lexogen UDI 12 nt Set B1 (UDI12B_0001-0096), 1 rxn/UDI |
| 156.384 | Lexogen UDI 12 nt Sets A1-A4 (UDI12A_0001-0384), 1 rxn/UDI |
UDI Add-on Kits (UDI plate and library amplification module) - Catalog Numbers
| Cat. No. | Product Name | UDI Add-on Kits (UDI plate and library amplification module) |
| 198.96 | Lexogen UDI 12 nt Unique Dual Indexing V2 Add-on Kit, Set A1 (UDI12A_0001-0096), 1 rxn/UDI |
| 199.96 | Lexogen UDI 12 nt Unique Dual Indexing V2 Add-on Kit, Set A2 (UDI12A_0097-0192), 1 rxn/UDI |
| 200.96 | Lexogen UDI 12 nt Unique Dual Indexing V2 Add-on Kit, Set A3 (UDI12A_0193-0288), 1 rxn/UDI |
| 201.96 | Lexogen UDI 12 nt Unique Dual Indexing V2 Add-on Kit, Set A4 (UDI12A_0289-0384), 1 rxn/UDI |
| 202.96 | Lexogen UDI 12 nt Unique Dual Indexing V2 Add-on Kit, Set B1 (UDI12B_0001-0096), 1 rxn/UDI |
| 203.384 | Lexogen UDI 12 nt Unique Dual Indexing V2 Add-on Kit, Sets A1-A4 (UDI12A_0001-0384), 1 rxn/UDI |
Buy from our Webstore
Need a web quote?
You can generate a web quote by Register or Login to your account. In the account settings please fill in your billing and shipping address. Add products to your cart, view cart and click the “Generate Quote” button. A quote in PDF format will be generated and ready to download. You can use this PDF document to place an order by sending it directly to sales@lexogen.com.
Web quoting is not available for countries served by our distributors. Please contact your local distributor for a quote.