TeloPrime Full-Length cDNA Amplification Kit V2 -
Effective capture of mature and intact mRNAs with exceptional 5' cap specificity
TeloPrime Full-Length cDNA Amplification Kit V2
TeloPrime Full-Length cDNA Amplification Kit V2 is an all-in-one protocol for generating full-length cDNA from total RNA and is based on Lexogen´s unique Cap-Dependent Linker Ligation (CDLL) and long reverse transcription (long RT) technology, which is highly selective for full-length RNA molecules that are both capped (5’ end) and polyadenylated (3’ end).
TeloPrime-generated full-length cDNA products can be used for various downstream applications such as NGS, RACE, cloning, microarray probes, and normalization. TeloPrime (Cat. No. 013) enables the detection and correct quantification of splice variants, and their true transcription start- and end-sites, in both short and long mRNA molecules. For further full-length or gene-specific PCRs, Lexogen offers a TeloPrime PCR Add-on Kit V2 (Cat. No. 018), containing separate PCR Forward and Reverse primers for alternative substitution with the gene-specific primer of interest.
Performance
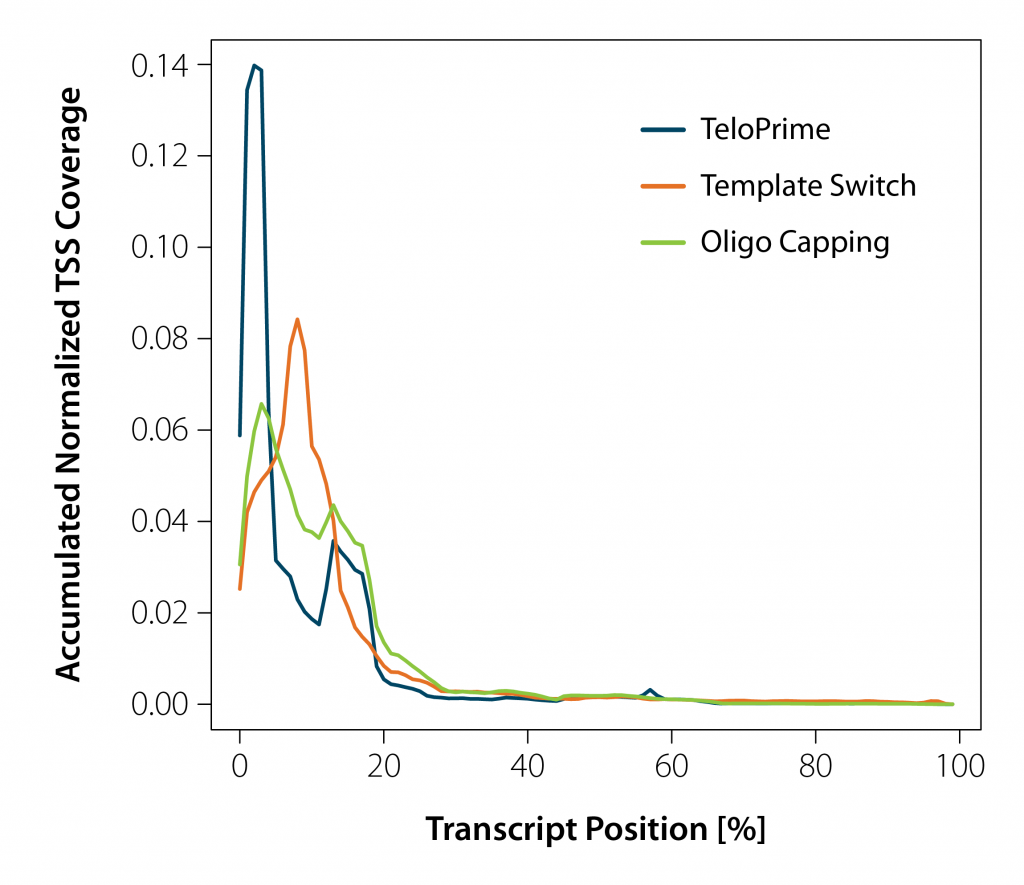
Full-length cDNA Synthesis of Mature and Intact mRNA
Workflow
10-18 hrs
1 hr 25 min
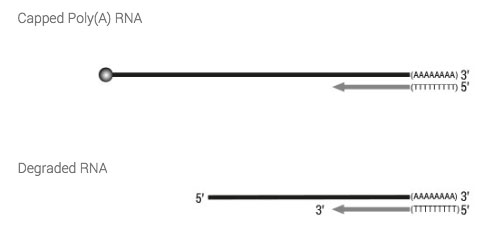
Firstly, full-length cDNA synthesis is initiated by oligodT primed long reverse transcription. This helps to preserve the complete RNA sequence information in the cDNA before a cap selection is carried out. In addition a more stable RNA/cDNA hybrid is created that is maintained throughout post RT purification. This double-stranded (ds) hybrid is also important for the specificity of the subsequent Cap-Dependent Linker Ligation reaction.
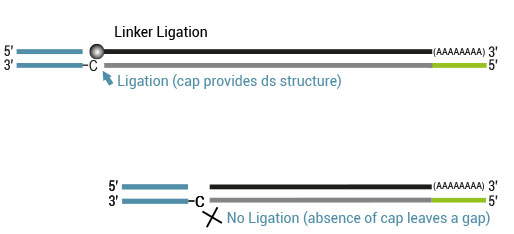
The double-stranded adapter with a 5’ C overhang allows for an atypical base-pairing with the inverted G of the cap structure. By using a ds-specific ligase, the ligation only takes place if the cap is present and if the RT has really reached the 5’ end of the mRNA. No ligation takes place if no cap is present e.g., in degraded RNA (low RIN) or if the RT has terminated prematurely because of secondary structures.
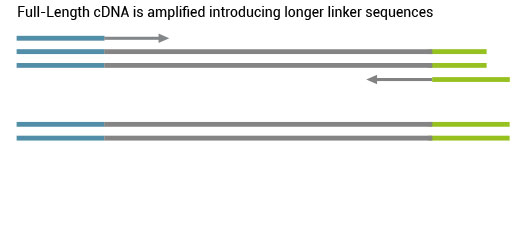
Based on this Cap-Dependent Linker Ligation only products with the 5’ tag get extended in the second-strand synthesis.
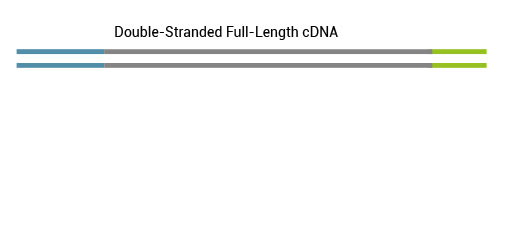
All remaining background is eliminated and the 5’ tagged full-length cDNA is converted into full-length ds cDNA.
Featured Publications
FAQ
Frequently Asked Questions
Access our frequently asked question (FAQ) resources via the buttons below.
Please also check our General Guidelines and FAQ resources!
How do you like the new online FAQ resource? Please share your feedback with us!
Downloads
Safety Data Sheet
If you need more information about our products, please contact us through support@lexogen.com or directly under +43 1 345 1212-41.
Ordering Information
| Cat. No. | Product Name |
| 013.08 | TeloPrime Full-Length cDNA Amplification Kit V2, 8 preps |
| 013.24 | TeloPrime Full-Length cDNA Amplification Kit V2, 24 preps |
| 018.16 | TeloPrime PCR Add-on Kit V2, 16 rxn |
First time user of TeloPrime?
First Time User? We’re excited to offer you an exclusive introductory offer.
Buy from our Webstore
Need a web quote?
You can generate a web quote by Register or Login to your account. In the account settings please fill in your billing and shipping address. Add products to your cart, view cart and click the “Generate Quote” button. A quote in PDF format will be generated and ready to download. You can use this PDF document to place an order by sending it directly to sales@lexogen.com.
Web quoting is not available for countries served by our distributors. Please contact your local distributor for a quote.