CORALL Total RNA-Seq V1
The CORALL Total RNA-Seq Library Prep Kit enables fast and cost-efficient generation of UMI labelled, stranded libraries for whole transcriptome analyses using Illumina® NGS platforms.
Dear Customer, CORALL V1 kits will be phased out soon. The last order date is October 31, 2023.
For more information, please get in touch with us at info@lexogen.com or sales@lexogen.com. Thank you!
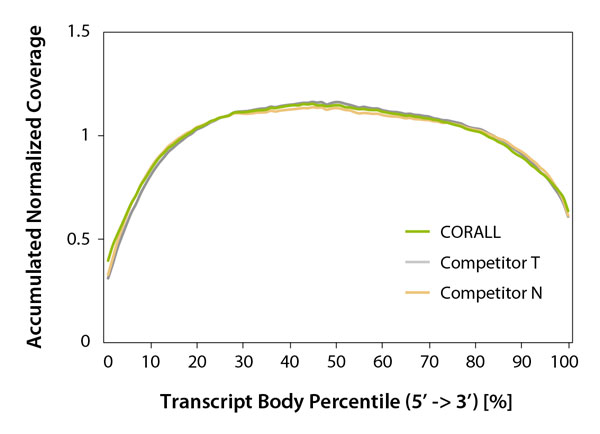
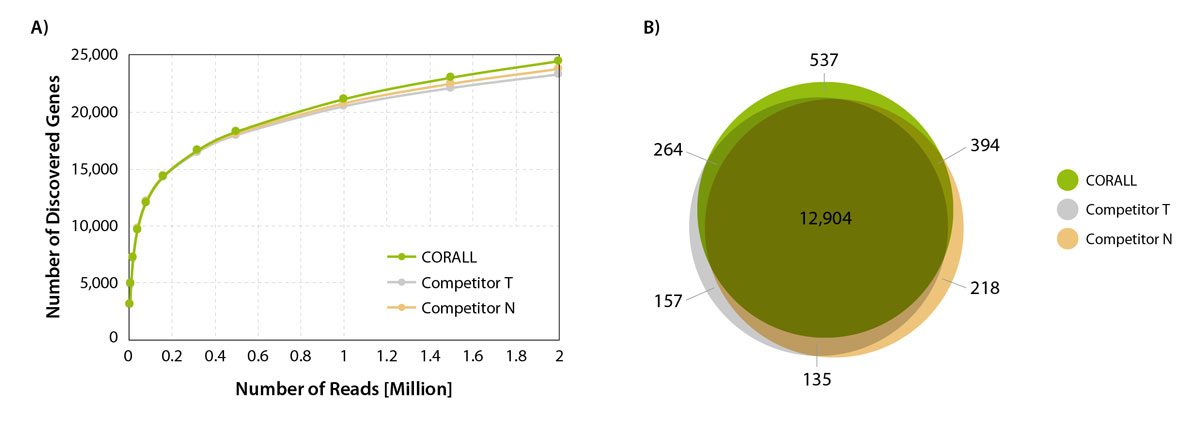
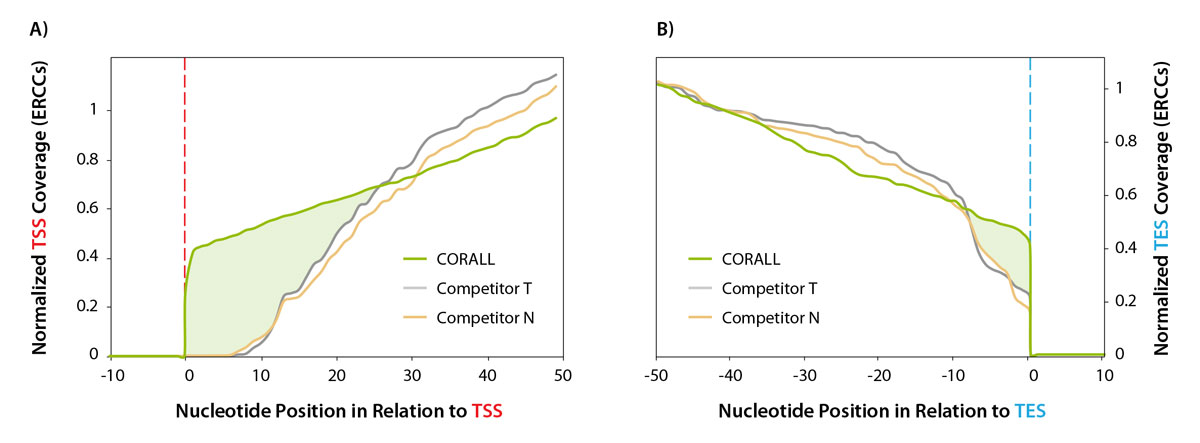
Performance
“We have successfully used the CORALL kit in our latest project to generate high quality sequencing libraries from low input material. With its fast and clear protocol we were able to generate the libraries in less than one day. We will surely use the CORALL kit again and recommend it for whole transcriptome analysis.”
“We used the new CORALL kit for performing transcriptome-analysis of CRISPR-modified cells in order to understand the consequences of deregulated epigenetic modifiers. In our hands the kit performance was highly satisfying in terms of data-quality and reproducibility across biological replicates. It furthermore convinced us with the ease of use, clarity of instructions, details in the manual, and handling of reagents.”
“CORALL kit has been key for our latest project. Despite having extremely heterogeneous samples, we were able to produce successful libraries from, literally, all our samples on the first attempt. Its perfect integration with Ribocop rRNA depletion kit and compatibility with SIRVs has made the project more straightforward than what we anticipated. Finally, as we required UMIs for demultiplexing, CORALL ticked all the boxes. We will use it again for our future projects.”
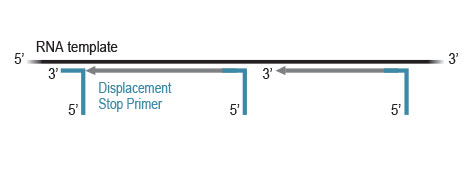
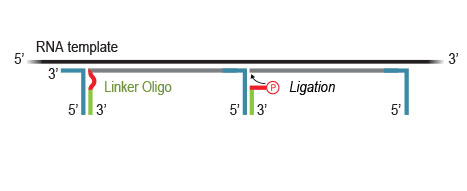
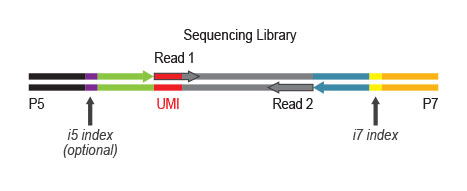
Workflow
FAQ
Frequently Asked Questions
Access our frequently asked question (FAQ) resources via the buttons below.
Please also check our General Guidelines and FAQ resources!
How do you like the new online FAQ resource? Please share your feedback with us!
Downloads
CORALL Total RNA-Seq Library Prep Kit for Illumina
![]() User Guide for CORALL Total RNA-Seq – update 31.03.2020
User Guide for CORALL Total RNA-Seq – update 31.03.2020
![]() User Guide for CORALL RNA-Seq with UDIs – update 28.03.2023
User Guide for CORALL RNA-Seq with UDIs – update 28.03.2023
![]() User Guide for Lexogen 12 nt Unique Dual Indexing Add-on Kit – update 25.01.2023
User Guide for Lexogen 12 nt Unique Dual Indexing Add-on Kit – update 25.01.2023
![]() Product Flyer – update 20.09.2019
Product Flyer – update 20.09.2019
![]() PCR Add-on Kit for Illumina User Guide – update 03.01.2023
PCR Add-on Kit for Illumina User Guide – update 03.01.2023
![]() Lexogen i5 6 nt Dual Indexing Add-on Kits (5001-5096) User Guide – update 03.01.2023
Lexogen i5 6 nt Dual Indexing Add-on Kits (5001-5096) User Guide – update 03.01.2023
![]() Lexogen UDI 12 nt Unique Dual Index Sequences – update 26.03.2021
Lexogen UDI 12 nt Unique Dual Index Sequences – update 26.03.2021
![]() Lexogen i7 and i5 Index Sequences – update 05.05.2020
Lexogen i7 and i5 Index Sequences – update 05.05.2020
Library Quantification File
Demultiplexing and Error Correction Tool – iDemux
Safety Data Sheet
If you need more information about our products, please contact us through support@lexogen.com or directly under +43 1 345 1212-41.
Buy from our Webstore
Need a web quote?
You can generate a web quote by Register or Login to your account. In the account settings please fill in your billing and shipping address. Add products to your cart, view cart and click the “Generate Quote” button. A quote in PDF format will be generated and ready to download. You can use this PDF document to place an order by sending it directly to sales@lexogen.com.
Web quoting is not available for countries served by our distributors. Please contact your local distributor for a quote.