RNA sequencing (RNA-Seq) has emerged as a cornerstone of cancer research and drug discovery, providing crucial insights for the development of novel therapies. By comprehensively profiling expression patterns, splice variants, and non-coding RNA effectors in solid and liquid tumors, RNA-Seq enables:
- Identification of novel compounds and molecules with anti-cancer activities and their mode-of-action,
- Investigation of the emergence and mechanisms of drug resistance,
- Discovery of biomarkers for diagnosis and prognosis, and
- to monitor treatment responses and disease progression.
In the following article, we will focus on the use of RNA-Seq and sophisticated data analyses aimed at overcoming drug resistance in various cancer models using the example of Forkhead box protein M1 (FOXM1) as target (Raghuwanshi et al., 2024).
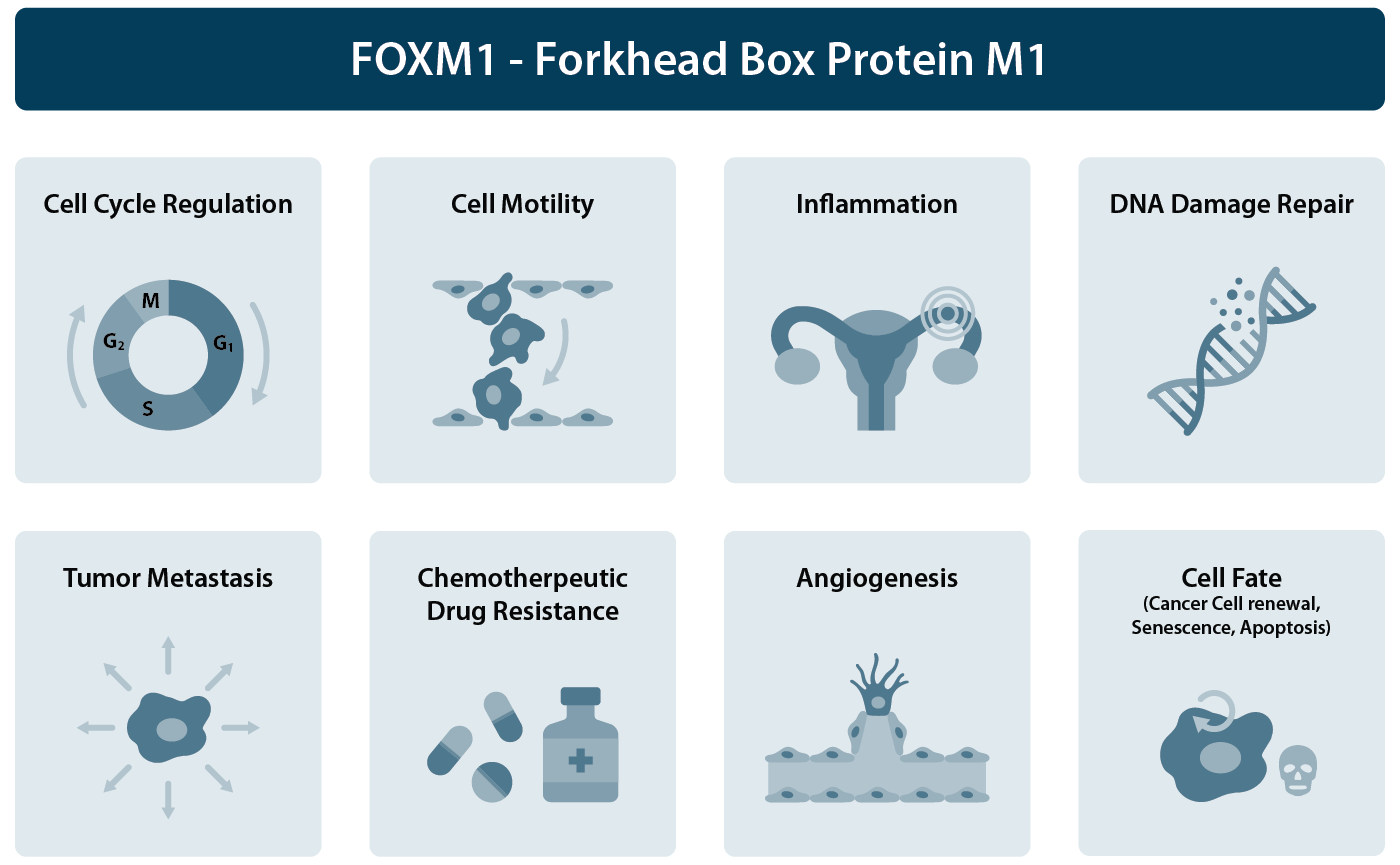
FOXM1 is a key transcription factor that plays a critical role in regulating essential cellular processes such as proliferation, the cell cycle, migration, DNA damage repair, redox signaling, and apoptosis (Fig. 1). Its overexpression is associated with the majority of human cancers (e.g., ovarian, breast, prostate, colorectal, and esophageal cancers). FOXM1 serves as a crucial regulator of tumor development impacting proliferation, survival, metastasis, inflammation, angiogenesis, and treatment resistance. Overexpression of FOXM1, for example due to crucial mutations or gene copy number amplification, elicits resistance to chemotherapy and is associated with poor patient outcomes.
FOXM1 drives tumor initiation, growth, and cancer progression
FOXM1 is expressed primarily in progenitor and regenerating tissue and is tightly controlled on the transcriptional, post-transcriptional, and post-translational level due to its critical role in proliferation (Liao et al., 2018, Anwar et al., 2024). Upon genetic alterations or lapses in said control, FOXM1 gets activated and/or overexpressed early on in cancer development and promotes cell cycle progression and proliferation, e.g., through c-Myc. Conversely, inhibition of FOXM1 in cancer cells reduces cell proliferation, migration, metastasis, and reduces the likelihood of resistance to cancer treatments appearing (Anwar et al., 2024). Several inhibitors of FOXM1 have been studied in various cancer models. They act on different regulatory levels from mRNA expression to translation, induce protein degradation or inhibit FOXM1 target binding. Known effectors range from small molecule inhibitors to non-coding RNAs, emphasizing the importance of FOXM1 regulation to improve patient outcomes.
Chemoresistance, whether inherent or acquired, remains a major obstacle in successful cancer treatment (Ramos et al., 2021, Vasan et al., 2019). While numerous molecular mechanisms contribute to the development of chemoresistance. FOXM1 consistently emerges as a key factor associated with reduced responsiveness to conventional chemotherapy across various tumor types (Fig. 1; Khan et al., 2024, Katzenellenbogen et al., 2023). Combination therapy has thus become paramount for cancer treatments counteracting, limitations in efficacy of single agent targeting drugs, especially when FOXM1 is involved.
Combination therapy targeting FOXM1 has the potential to overcome chemoresistance in various human cancers
In 2021, researchers from the University of Illinois at Chicago and the College of Medicine at the University of Pittsburgh used a network-based screening approach of differentially expressed gene signatures to identify small molecule compounds inhibiting FOXM1-associated pathways. This analysis revealed STL427944 as a potential new inhibitor of FOXM1 with the ability to sensitize chemoresistant cancer cells towards three categories of widely used anticancer drugs (Chesnokov et al., 2021).
In a follow-up study from 2024, the authors investigate STL001, a first-generation modification drug of the previously reported FOXM1 inhibitor STL427944. While the modified version maintains the mode-of-action of STL427944 it could be shown to be up to ~50 times more potent in a variety of solid cancers (Raghuwanshi et al., 2024).
Whole transcriptome sequencing, utilizing RiboCop rRNA Depletion and CORALL RNA-Seq, confirmed the mechanism of action for STL001. By comparing STL001-treated cells with those treated with its predecessor compound STL427944 and FOXM1-knockdown cells across multiple cancer models, the authors not only identified known FOXM1-regulated pathways but uncovered novel activities. STL001 effectively suppresses FOXM1 by inhibiting its transcriptional activity. This leads to decreased expression of FOXM1-regulated genes involved in cell proliferation, survival, and resistance to chemotherapy. Importantly, treatment with STL001 significantly enhanced the sensitivity of various cancer cell lines (lung, ovarian, colorectal) to multiple chemotherapeutic drugs (e.g., cisplatin, gemcitabine, and doxorubicin), suggesting its tremendous potential as a promising partner drug for combination therapy to overcome resistance in solid tumors.
Omics Playground reveals striking correlations between STL001 and mTOR inhibitors and identifies tetrandrine as a high-potential partner drug
Further analyses of the publicly available dataset from Raghuwanshi et al. using BigOmics Analytics’ Omics Playground platform confirmed the authors conclusions and revealed additional insights. Strikingly, STL001 and mTOR/PI3K inhibitors share similar expression profiles – likely due to the interconnected roles of FOXM1 and mTOR in cell growth, proliferation, and survival. Both pathways interact in cell cycle regulation, DNA repair, and metabolism, often targeting common downstream processes (e.g., mitotic checkpoints, DNA repair). Since both also enhance apoptosis, combining STL001 with mTOR inhibitors could create synergistic anticancer effects. In addition, the platform identified tetrandrine – a plant alkaloid with anti-cancer properties, as a potential partner drug. Tetrandrine shows a similar gene expression profile to STL001 albeit through acting via different pathways. Hence, combining these compounds could be particularly effective and offer novel possibilities for combination strategies with greater therapeutic efficacy.
For more detailed data analyses insights, read the full blog: Advancing Cancer Therapies: Synergizing mTOR Inhibitors with STL001’s Enhanced Modes of Action and Tetrandrine Potential.
Beyond analysis: Investigating novel combination strategies to overcome drug resistance human cancer
Drug response profiling to assess optimal dose-response of drug combinations
RNA-Seq experiments are extremely useful to generate dose-dependent expression profiles for single drugs and compound combinations. By comparing the expression profiles pre- and post combination treatment across various concentrations, researchers gain comprehensive insights on how the compounds affect gene regulatory networks, at which concentration the combination is most potent, and whether drug concentrations are indicative of physiological administration ranges. Ideal combinations show a greater-than-additive effect and synergy can be confirmed through analyses on gene expression and pathway level.
Dose-response experiments are typically done using cancer-related cell culture models. In this case, it would be important to assess the effects of combination therapies and concentration ranges in various models. This allows researchers to draw biologically valid conclusions for various cancer types when targeting master regulators associated with a majority of human cancers. Dose-dependent RNA-Seq analysis can confirm in silico predictions for partner drugs and even identify additional targets which can be hit by combination approaches or become affected at different concentrations. In addition, time-resolved expression analysis using SLAMseq allows to unravel primary and secondary effects of fast-acting drugs, response sequences, and interdependencies of drug combinations. Explore our previous blog for a comprehensive overview on RNA-Seq in Drug Discovery and Development and the importance of time-resolved RNA sequencing.
Of note, it is always worth to consult with experts in bioinformatics, cancer biology, and next-generation sequencing (NGS) to design and analyze RNA-seq experiments for drug combination studies effectively. As the field is rapidly advancing, service providers allow easy access to state-of-the-art wet lab and analyses methods to ensure success through a data-driven approach.
Circumventing drug resistance in cancer treatments
Combination therapy aims to circumvent drug resistance. Hence, following the confirmation of the potential efficacy of a drug combination, it is crucial to understand which resistance mechanisms can be overcome and for which the approach is not suitable. To this end, researchers use RNA-Seq and reporter assays in resistance models to identify for which conditions the previously determined dose is effective. By assessing the mode-of-action in correlation with the resistance mechanism and interrogating suitable biomarkers, the treatments potential can be further elucidated. In the long-term, this information aids patient stratification and the development of different treatment options tailored to the specific genetic make-up and potential risks to develop resistances.
Biomarkers discovery
The identification of expression signatures indicative of the response to the treatment plays a pivotal role in biomarker discovery. These signatures need to be further validated through biological systems from cell culture to animal models and patients to generate tests that can stratify patient groups for tailored, precise treatment according to their specific cancer phenotype.
Key considerations for experiments to explore combination therapy and validate predicted partner drugs
Careful experimental design and stringent data analysis are crucial for obtaining meaningful results from RNA-seq experiments in drug combination studies.
Key considerations for designing RNA-Seq experiments for drug discovery and drug combinations:
- Ensure sufficient sample size to increase statistical power and include biological replicates for treatment and control conditions. Consult with Bioinformaticians and data teams to optimize the experimental setup.
- Include suitable controls: untreated, diluent controls, vector controls, scrambled sequences when using oligomers, and artificial spike-in controls for in-process and quality control or normalization.
- When performing large-scale or long-term experiments, select plate layouts that allow correction for batch effects throughout the experiment.
- Consider dose-response experiments to determine optimal drug concentrations for single drug administrations and combinations.
- Include multiple time points and/or perform time-resolved RNA-Seq to capture dynamic changes in gene expression and determine primary and secondary drug effects.
- Randomize sample to minimize bias where possible.
Overall, integrating RNA-seq data with additional experimental datasets, such as cell viability assays, flow cytometry, ‘omics’ data etc. is required to validate the observed effects before moving to more sophisticated systems like organoids and animal models. The latter allows researchers to add additional layers, e.g., tumor heterogeneity, for a data-driven approach to treatment design when investigation novel combination treatment strategies.
Summary and key findings
RNA-Seq is crucial for cancer research and drug discovery, providing insights into gene expression and aiding in the development of new therapies. This article focuses on STL001, a newly identified inhibitor of FOXM1 – a transcription factor linked to cancer progression and drug resistance.
Key findings:
- Whole transcriptome RNA-Seq was used to elucidate the mode-of-action and to confirm potential synergies of STL001 with other drugs known to be more effective upon FOXM1 inhibition.
- Further data analyses using Omics Playground with the public data set suggest additional, so far unexplored potential synergies between STL001 and mTOR inhibitors, as well as tetrandrine. Read the blog: Advancing Cancer Therapies: Synergizing mTOR Inhibitors with STL001's Enhanced Modes of Action and Tetrandrine Potential.
- These insights can be further explored to exploit the versatility of STL001 and its use in combination therapies to combat cancer drug resistance.
Follow-up RNA-Seq experiments can be designed, following a set of key considerations, to further investigate optimal drug combinations with STL001, understand its applicability in context of various resistance mechanisms, and discover biomarkers indicative of treatment responses. End-to-end solutions with state-of-the-art wet lab and data analyses workflows can accelerate the development of more effective cancer therapies.
About Lexogen
Lexogen is a leading provider of reagents and NGS services, specialized in RNA sequencing. Lexogen NGS Services offers a comprehensive range of sequencing services, including RNA-Seq, single-cell sequencing and DNA-Seq. Lexogen’s expertise in RNA biology and commitment to quality and efficiency empowers our clients’ discoveries. Project-tailored services and rigorous quality control ensure accurate and reliable results even for the most demanding applications. Consult with us to accelerate your insights!
About BigOmics Analytics
BigOmics Analytics is a Swiss company that expedites tertiary analysis, by providing a centralized platform solution for scientists to efficiently scale their omics data analysis and obtain reproducible results. Through its collaborative analysis platform, Omics Playground, researchers can interactively share data and insights, fostering efficient collaboration. For more information, visit BigOmics’ website.
Omics Playground
Omics Playground provides highly interactive and user-friendly tools for understanding RNA-seq and proteomics data. The platform helps biotech, pharma and academic teams to reduce the time spent on data analysis and free up more time for discovery. With more than 18 analysis modules and 150 interactive plots, Omics Playground uses best-in-class methods and algorithms for each analysis step, ensuring robust and reproducible results. To further enhance its capabilities, BigOmics is expanding support to metabolomics and multi-omics analysis, broadening the platform’s reach across diverse omics disciplines.
Read the publications
Dive deeper into the discovery of STL001 below.
Abstract
FOXM1 transcription factor is an oncogene and a master regulator of chemoresistance in multiple cancers. Pharmacological inhibition of FOXM1 is a promising approach but has proven to be challenging. We performed a network-centric transcriptomic analysis to identify a novel compound STL427944 that selectively suppresses FOXM1 by inducing the relocalization of nuclear FOXM1 protein to the cytoplasm and promoting its subsequent degradation by autophagosomes. Human cancer cells treated with STL427944 exhibit increased sensitivity to cytotoxic effects of conventional chemotherapeutic treatments (platinum-based agents, 5-fluorouracil, and taxanes). RNA-seq analysis of STL427944-induced gene expression changes revealed prominent suppression of gene signatures characteristic for FOXM1 and its downstream targets but no significant changes in other important regulatory pathways, thereby suggesting high selectivity of STL427944 toward the FOXM1 pathway. Collectively, the novel autophagy-dependent mode of FOXM1 suppression by STL427944 validates a unique pathway to overcome tumor chemoresistance and improve the efficacy of treatment with conventional cancer drugs.
Abstract
Forkhead box protein M1 (FOXM1) is often overexpressed in human cancers and strongly associated with therapy resistance and less good patient survival. The chemotherapy options for patients with the most aggressive types of solid cancers remain very limited because of the acquired drug resistance, making the therapy less effective. NPM1 mutation through the inactivation of FOXM1 via FOXM1 relocalization to the cytoplasm confers more favorable treatment outcomes for AML patients, confirming FOXM1 as a crucial target to overcome drug resistance. Pharmacological inhibition of FOXM1 could be a promising approach to sensitize therapy-resistant cancers. Here, we explore a novel FOXM1 inhibitor STL001, a first-generation modification drug of our previously reported FOXM1 inhibitor STL427944. STL001 preserves the mode of action of the STL427944; however, STL001 is up to 50 times more efficient in reducing FOXM1 activity in a variety of solid cancers. The most conventional cancer therapies studied here induce FOXM1 overexpression in solid cancers. The therapy-induced FOXM1 overexpression may explain the failure or reduced efficacy of these drugs in cancer patients. Interestingly, STL001 increased the sensitivity of cancer cells to conventional cancer therapies by suppressing both the high-endogenous and drug-induced FOXM1. Notably, STL001 does not provide further sensitization to FOXM1-KD cancer cells, suggesting that the sensitization effect is conveyed specifically through FOXM1 suppression. RNA-seq and gene set enrichment studies revealed prominent suppression of FOXM1-dependent pathways and gene ontologies. Also, gene regulation by STL001 showed extensive overlap with FOXM1-KD, suggesting a high selectivity of STL001 toward the FOXM1 regulatory network. A completely new activity of FOXM1, mediated through steroid/cholesterol biosynthetic process and protein secretion in cancer cells was also detected. Collectively, STL001 offers intriguing translational opportunities as combination therapies targeting FOXM1 activity in a variety of human cancers driven by FOXM1.
Explore additional data analyses insights: Advancing Cancer Therapies: Synergizing mTOR Inhibitors with STL001’s Enhanced Modes of Action and Tetrandrine Potential.
References
Anwar, S., Zafar, M., Hussain, M.A., Iqbal, N., Ali, A., Sadaf, Kaur, S., Najm, M.Z., and Kausar, M.A. (2024) Unravelling the therapeutic potential of forkhead box proteins in breast cancer: An update (Review). Oncol Rep. 52(1):92. DOI: 10.3892/or.2024.8751. Epub. PMID: 38847267; PMCID: PMC11177173.
Chesnokov, M.S., Halasi, M., Borhani, S., Arbieva, Z., Shah, B.N., Oerlemans, R., Khan, I., Camacho, C.J., and Gartel, A.L. (2021) Novel FOXM1 inhibitor identified via gene network analysis induces autophagic FOXM1 degradation to overcome chemoresistance of human cancer cells. Cell Death Dis 12, 704. DOI: 10.1038/s41419-021-03978-0
Kalathil, D., John, S., and Nair, A.S. (2021) FOXM1 and Cancer: Faulty Cellular Signaling Derails Homeostasis. Front Oncol. 10:626836. DOI: 10.3389/fonc.2020.626836. PMID: 33680951; PMCID: PMC7927600.
Katzenellenbogen, B.S., Guillen, V.S. and Katzenellenbogen, J.A. (2023) Targeting the oncogenic transcription factor FOXM1 to improve outcomes in all subtypes of breast cancer. Breast Cancer Res 25, 76. DOI: 10.1186/s13058-023-01675-8
Khan, S.U., Fatima, K., Aisha, S. and Malik, F. (2024) Unveiling the mechanisms and challenges of cancer drug resistance. Cell Commun Signal 22, 109. DOI: 10.1186/s12964-023-01302-1
Liao, GB., Li, XZ., Zeng, S., Liu, E., Yang, SM., Yang, L., Hu, CJ., and Bai, JY. (2018) Regulation of the master regulator FOXM1 in cancer. Cell Commun Signal 16, 57. DOI: 10.1186/s12964-018-0266-6
Raghuwanshi, S., Zhang, X., Arbieva, Z., Khan, I., Mohammed, H., Wang, Z., Domling, A., Camacho, C. J., and Gartel, A. L. (2024) Novel FOXM1 inhibitor STL001 sensitizes human cancers to a broad-spectrum of cancer therapies. Cell Death Discov. 10, 211. DOI: 10.1038/s41420-024-01929-0
Ramos, A., Sadeghi, S., and Tabatabaeian, H. (2021) Battling Chemoresistance in Cancer: Root Causes and Strategies to Uproot Them. Int J Mol Sci. 22(17):9451. DOI: 10.3390/ijms22179451. PMID: 34502361; PMCID: PMC8430957.
Vasan, N., Baselga, J., and Hyman, D.M. (2019) A view on drug resistance in cancer. Nature 575, 299–309. DOI: 10.1038/s41586-019-1730-1
Written by Dr. Yvonne Göpel







