RNA sequencing (RNA-Seq) has become an indispensable tool for cancer researchers. Unlike traditional methods, RNA-Seq provides a dynamic snapshot of the complete transcriptome, revealing not just the presence of specific genes but also their expression level at a given time, e.g., during disease progression or treatment. While DNA sequencing illuminates the genetic landscape of tumors, RNA-Seq not only reveals the functional consequences of genetic alterations, but also unravels underlying regulatory mechanisms and can identify aberrant transcript isoforms linked to the disease.
How does RNA-Seq compare to classical approaches to study cancer?
Cancer research employs diverse methods, ranging from histology and imaging to visualize lesions, to flow cytometry for immune cell involvement. Since cancer fundamentally involves genomic changes, cancer genome sequencing is crucial for understanding genetic variants. Transcriptional changes, i.e., how genes are expressed in cancer versus healthy tissue or after treatment are typically assessed with techniques like RT-qPCR, microarrays, and RNA-Seq.
RT-qPCR is a low-throughput method suitable for detecting and quantifying a small panel of target mRNA molecules. Microarrays, while transformative for cancer research by providing more comprehensive gene expression data, are limited by their reliance on prior knowledge of gene sequences, meaning they cannot identify novel gene expression patterns. RNA-Seq, on the other hand, is a high-throughput technology that can detect novel transcripts, gene fusions, single nucleotide variants, indels, and other subtle changes that microarrays cannot. It boasts a much larger dynamic range for quantification and higher sensitivity, enabling the detection of a greater percentage of differentially expressed genes, even those with low abundance. Furthermore, advancements in RNA-Seq now permit transcriptomic investigation at single-cell resolution, far surpassing the sensitivity of traditional molecular biology methods.
Why is RNA-Seq so crucial in cancer research?
Cancer cells are typically characterized by genomic instability and the accumulation of a plethora of mutations. RNA-Seq allows us to focus on the biologically relevant changes – the specific gene alterations that drive tumor progression, metastasis, recurrence, and resistance to treatment. By interrogating only the expressed transcripts, RNA-Seq effectively “sorts the needles in the haystack,” enabling researchers to identify the key drivers of malignancy. Combining DNA and RNA sequencing additionally allows to classify mutation severity and gain a holistic view of the tumor biology.
How is RNA-Seq contributing to our fundamental understanding of cancer biology? A few examples.
1. Decoding Gene Function and Pathways:
Differential gene expression analysis, a cornerstone of RNA-Seq, reveals how gene activity fluctuates across different cancer states, such as varying stages of progression, responses to therapy, or differences between tumor subtypes. This analysis uncovers crucial molecular mechanisms and regulatory pathways that contribute to cancer development and progression. Both 3′ mRNA-Seq, which focuses on the 3′ end of transcripts for efficient gene quantification, and whole transcriptome RNA-Seq, which captures the full length of transcripts including isoforms and non-coding RNAs, are widely used for comprehensive transcriptional profiling in cancer research, each offering unique insights.
2. Discovering Cancer Biomarkers:
RNA-Seq is instrumental in identifying diverse cancer biomarkers, including those indicating tumor progression, predicting recurrence, and forecasting treatment response. These biomarkers can be used for early detection, personalized treatment strategies, and patient stratification. Small RNAs (miRNAs) and various non-coding RNAs (such as lncRNAs and circRNAs) play diverse regulatory roles in cancer development and progression which can be identified by RNA-Seq. Small RNAs and other non-coding RNAs are continuously recognized as potent biomarkers. In addition, gene fusions which often drive oncogenesis, are detected by RNA-Seq. Targeted approaches enhance sensitivity for detecting specific fusion transcripts.
3. Mapping Tumor Heterogeneity and Unraveling Drug Resistance:
Single-cell RNA-Seq (scRNA-Seq) and spatial RNA-Seq provide unprecedented resolution, revealing gene expression patterns in individual tumor cells and can preserve information on morphology and localization. These technologies have helped elucidate the vast heterogeneity within tumors which is crucial for understanding cancer progression and drug resistance. On a global scale, RNA-Seq can help identify mutations, gene expression changes, and pathway alterations that drive cancer drug resistance, enabling the development of strategies to overcome resistance. Check out our recent blog for more information on overcoming chemoresistance with multimodal combination therapy.
How RNA-Seq revolutionizes precision medicine and cancer treatment for better patient outcomes - a case study
Immune checkpoint inhibitors in cancer therapy
One theme in immunotherapy is the use of immune checkpoint inhibitors, short ICIs. Immune checkpoints are part of the natural immune system – they are decision points for the initiation of an immune reaction, e.g., against foreign cells such as pathogens or cancer cells which are growing unnaturally. Upon activation, the immune system can attack cancer cells, e.g., through T-cells while preserving normal cells. However, cancer cells have evolved to evade immune reactions fostering unchecked tumor growth and cancer progression. Immune checkpoint inhibitors, such as Pembrolizumab, an anti-PD-1 inhibitor, are used as therapeutics to reverse the cancer evasion and force an immune response (Fig. 2).
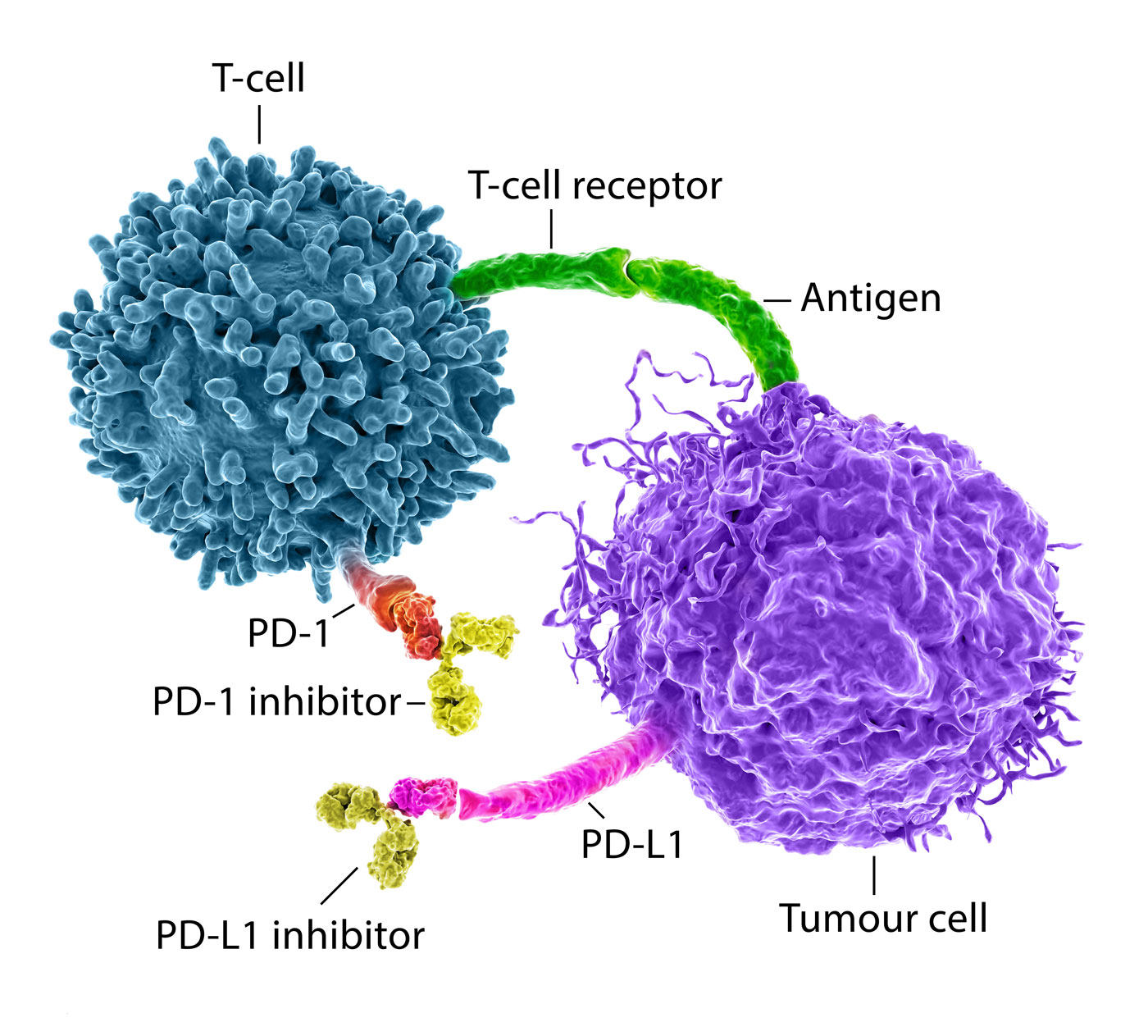
Unfortunately, ICIs are not effective for many patients and ICI therapies are often supplemented with chemotherapy which may cause severe side effects. For example, immunohistochemistry tests are used to detect the presence of PD-L1 and determine patients who may respond to ICI monotherapy. The test uses a scoring system that correlates to the protein amounts and aims to predict responders. Based on the cancer type and considered therapy, the scoring system for PD-L1 differs, further complicating classification for physicians. While widely administered, the test itself may classify some patients as undetermined or falsely identify them as non-responders. Because of these limitations, patients are often treated with aggressive chemotherapy in addition to ICI therapy and many experience a range of side effects. Stringent stratification and sensitive detection are prerequisites for effective patient care and a must if harsh therapies should be avoided to overall improve the quality of life for patients during treatment all the while efficiently fighting cancer (Marei et al., 2023).
The potential of RNA-Seq for cancer patient testing and stratification
A recently developed test termed OncoPrism™ uses RNA sequencing and machine learning to stratify patients into treatment groups to improve outcomes for patients with recurrent/metastatic head and neck squamous cell carcinoma (Flanagan et al., 2023). OncoPrism is an RNA-based multi-analyte biomarker test that predicts the response to ICI in cancer patients by using health expression models. Gene expression patterns can be more indicative of a patients’ predicted response as compared to single-analyte immunohistochemistry tests and often provide a more sensitive read-out.
In 2024, a multicenter study termed PREDAPT (PREDicting immunotherapy efficacy from Analysis of Pre-treatment Tumor biopsies) was conducted to validate the OncoPrism-HNSCC assay across 17 US healthcare systems. The study used a 99-patient training cohort and two independent validation cohorts of 62 and 50 patients (Flanagan et al., 2024). The returned OncoPrism Score stratifies patients into low, medium, and high likelihood of disease control with more than threefold higher specificity compared to PD-L1 testing and approximately fourfold higher sensitivity than tumor mutational burden for predicting disease control.
The prerequisite to develop reliable and sensitive tests for patients and the success of clinical validations is the underlying robustness and analytical performance of the technology and wet lab workflow used to generate the data (Hiken et al., 2025). High quality data is strongly associated with reproducible and consistent sample preps that can use even suboptimal input materials. Consistent performance within and across batches or processing units are key for conclusive expression pattern measurements. Systematic variations are known to occur in biological data sets. However, they typically arise from non-biological factors and are the result of experimental processing, such as differences in equipment, reagents, or operators. These variations need to be characterized and controlled as they can obscure true biological differences result in inaccurate conclusions. Streamlined and robust sample preps with fewer protocol steps and internal controls increase consistency and reproducibility.
The RNA-Seq library preparation used to generate the health expression models for OncoPrism predictions is QuantSeq – a renowned 3′ mRNA-Seq method that was first released in 2014 and pioneered the advance of expression profiling through RNA sequencing (Moll et al. 2014). RNA-Seq libraries are generated in only 5 steps without the need for prior depletion (Fig. 3). The workflow uses in-prep poly(A) selection and generates strand-specific libraries at the 3′ end of polyadenylated transcripts – making it suitable for mRNA profiling even from low-quality FFPE samples, which are often a challenge for cancer research and diagnostics. For more information on FFPE samples and RNA-Seq, also check our recent blog articles on FFPE transcriptomics.
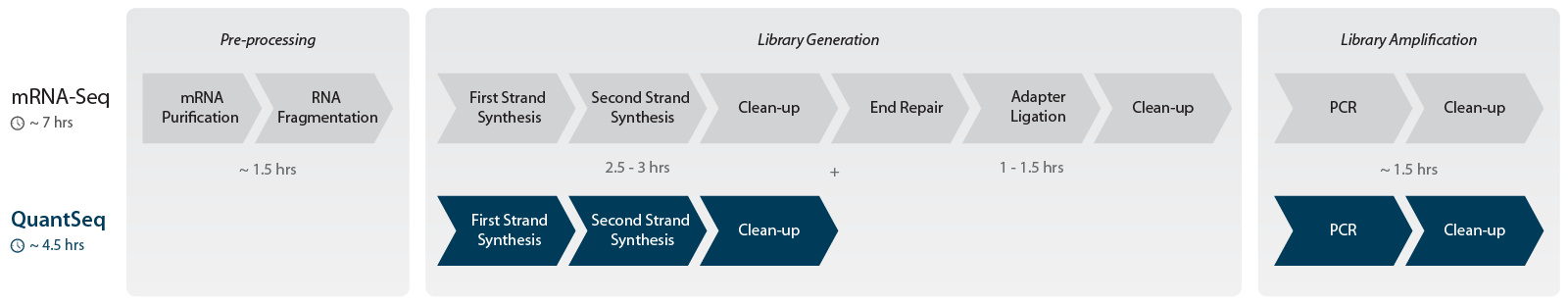
Interested to learn more about OncoPrism and QuantSeq?
Watch the webinar with Kevin Flanagan, PhD – Director of Translational Science at Cofactor Genomics.
The following publications summarize the development and validation of OncoPrism™, a biomarker classifier to predict the response of cancer patients to immune checkpoint therapy.
This study aimed to develop a new biomarker classifier, OncoPrism-HNSCC, to predict disease control in response to anti-PD-1 therapy in patients with recurrent or metastatic head and neck squamous-cell carcinoma (RM-HNSCC). The OncoPrism Score ranges from 0 to 100 and correlates with disease control and overall survival in patients treated with anti-PD-1 monotherapy. Statistics were conducted using R software tools, and the biomarker was internally validated. Pre-treatment tumor samples from 172 patients were collected, with 103 samples meeting inclusion criteria. RNA sequencing and biomarker training resulted in the selection of 62 immunomodulatory features for the final model. The OncoPrism Score predicted disease control with high sensitivity and specificity.
Abstract
Purpose
Anti-PD-1 therapy provides clinical benefit in 40–50% of patients with relapsed and/or metastatic head and neck squamous cell carcinoma (RM-HNSCC). Selection of anti- PD-1 therapy is typically based on patient PD-L1 immunohistochemistry (IHC) which has low specificity for predicting disease control. Therefore, there is a critical need for a clinical biomarker that will predict clinical benefit to anti-PD-1 treatment with high specificity.
Methods
Clinical treatment and outcomes data for 103 RM-HNSCC patients were paired with RNA-sequencing data from formalin-fixed patient samples. Using logistic regression methods, we developed a novel biomarker classifier based on expression patterns in the tumor immune microenvironment to predict disease control with monotherapy PD-1 inhibitors (pembrolizumab and nivolumab). The performance of the biomarker was internally validated using out-of-bag methods.
Results
The biomarker significantly predicted disease control (65% in predicted non-progressors vs. 17% in predicted progressors, p < 0.001) and was significantly correlated with overall survival (OS; p = 0.004). In addition, the biomarker outperformed PD-L1 IHC across numerous metrics including sensitivity (0.79 vs 0.64, respectively; p = 0.005) and specificity (0.70 vs 0.61, respectively; p = 0.009).
Conclusion
This novel assay uses tumor immune microenvironment expression data to predict disease control and OS with high sensitivity and specificity in patients with RM-HNSCC treated with anti-PD-1 monotherapy.
A multicenter study aimed to validate the RNA-based OncoPrism-HNSCC assay, designed to predict disease control in patients undergoing anti-PD-1 treatment. The study involved a total of 211 samples, divided into a training cohort of 99 patients and two validation cohorts of 62 and 50 patients, respectively. The assay classifies patients into three groups based on their likelihood of disease control: low, medium, and high, with corresponding OncoPrism Scores generating from RNA-sequencing data.
Abstract
Background
Despite advances in cancer care and detection, >65% of patients with squamous cell cancer of the head and neck (HNSCC) will develop recurrent and/or metastatic disease. The prognosis for these patients is poor with a 5-year overall survival of 39%. Recent treatment advances in immunotherapy, including immune checkpoint inhibitors like pembrolizumab and nivolumab, have resulted in clinical benefit in a subset of patients. There is a critical clinical need to identify patients who benefit from these antiprogrammed cell death protein 1 (anti-PD-1) immune checkpoint inhibitors.
Methods
Here, we report findings from a multicenter observational study, PREDicting immunotherapy efficacy from Analysis of Pre-treatment Tumor biopsies (PREDAPT), conducted across 17 US healthcare systems. PREDAPT aimed to validate OncoPrism-HNSCC, a clinical biomarker assay predictive of disease control in patients with recurrent or metastatic HNSCC treated with anti-PD-1 immune checkpoint inhibitors as a single agent (monotherapy) and in combination with chemotherapy (chemo-immunotherapy). The test used RNA-sequencing data and machine learning models to score each patient and place them into groups of low, medium, or high.
Results
The OncoPrism-HNSCC prediction significantly correlated with disease control in both the monotherapy cohort (n=62, p=0.004) and the chemo-immunotherapy cohort (n=50, p=0.01). OncoPrism-HNSCC also significantly predicted progression-free survival in both cohorts (p=0.015 and p=0.037, respectively). OncoPrism-HNSCC had more than threefold higher specificity than programmed death-ligand 1 combined positive score and nearly fourfold higher sensitivity than tumor mutational burden for predicting disease control.
Conclusions
Here, we demonstrate the clinical validity of the OncoPrism-HNSCC assay in identifying patients with disease control in response to anti-PD-1 immune checkpoint inhibitors.
Abstract
Background
While immune checkpoint inhibitor (ICI) therapies can significantly improve outcomes for patients with recurrent/metastatic head and neck squamous cell carcinoma (RM-HNSCC), only about 15–20% benefit from such treatments. Clinical tests that guide the use of ICIs are therefore critically needed. OncoPrism-HNSCC was developed to address this need. The assay combines next generation RNA sequencing-based immunomodulatory gene expression signatures with machine learning algorithms to generate an OncoPrism score that classifies patients as having low, medium, or high likelihood of disease control in response to ICI treatment. Also, OncoPrism-HNSCC leverages the same FFPE patient tumor RNA used for ICI response prediction to identify rare cases where oncogenic rearrangements in NTRK1/2/3 or ALK genes may occur, and which may indicate the use of potentially highly effective targeted therapies. The clinical performance of OncoPrism-HNSCC has been validated. Here, we report its analytical performance in the presence of potentially confounding sources of variation.
Methods
The assay’s analytical sensitivity was assessed by varying RNA input quantity and quality, observing the effect on ICI response prediction scores. Analytical specificity was tested by spiking increasing percentages of genomic DNA into input RNA. Intra-assay and inter-assay precision were evaluated, and the analytical sensitivity, specificity, and precision of gene fusion detection were assessed. Concordance with orthogonal methods of gene fusion detection was tested on 67 FFPE clinical samples.
Results
Varying RNA inputs as low as four-fold below the nominal input amount had little effect on ICI response prediction scores. RNA quality levels below the test threshold had no significant effect. Genomic DNA spike-ins up to 30% had only a small effect on scores. The pooled standard deviation for multiple operators, reagent lots, batches, and sequencers yielded an overall variance represented by just 0.87% of the score range of the test (0–100). NTRK and ALK gene fusion detection was 100% concordant with orthogonal methods.
Conclusions
Robust and reliable analytical performance of the OncoPrism-HNSCC assay supports its clinical use, even in the presence of variation typically encountered in the laboratory setting.
Summary
RNA-Seq is an emerging tool in cancer research and diagnotics, providing a dynamic and comprehensive view of gene expression offering significant advantages over RT-qPCR and microarrays due to the ability to detect novel transcripts, higher sensitivity, single-cell resolution, and upscaling potential.
In precision medicine, RNA-Seq is revolutionizing cancer patient stratification, particularly for immunotherapies. By providing more sensitive and specific gene expression profiles than conventional tests (like PD-L1 IHC), RNA-Seq-based assays, such as OncoPrism™, enable better patient selection for treatments and contribute to improved patient outcomes.
References
Flanagan, K.C., Earls, J., Schillebeeckx, I., Hiken, J., Wellinghoff, R.L., LaFranzo, N.A., Bradley, Z.S., Babbitt, J., Westra, W.H., Hsu, R., Nadauld, L., Mcleod, H., Firth, S.D., Sharp, B., Fuller, J., Vavinskaya, V., Sutton, L., Deichaite, I., Bailey, S.D., Sandulache, V.C., Rendo, M.J., Macdonald, O.K., Welaya, K., Wade, J.L. 3rd, Pippas, A.W., Slim, J., Bank, B., Saccaro, S.J., Sui, X., Akhtar, A., Balaraman, S., Kossman, S.E., Sonnier, S.A., Shenkenberg, T.D., Alexander, W.L., Price, K.A., Bane, C.L., Ley, J., Messina, D.N., Glasscock, J.I., Cohen, E.E.W., Adkins, D.R., and Duncavage, E.J. (2023) Multidimensional biomarker predicts disease control in response to immunotherapy in recurrent or metastatic head and neck squamous-cell carcinoma. J Cancer Res Clin Oncol. 149:14125-14136. DOI: 10.1007/s00432-023-05205-z. PMID: 37552307; PMCID: PMC10590294.
Flanagan, K.C., Earls, J., Hiken, J., Wellinghoff, R.L., Ponder, M.M., McLeod, H.L., Westra, W.H., Vavinskaya, V., Sutton, L., Deichaite, I., Macdonald, O.K., Welaya, K., Wade, J. 3rd, Azzi, G., Pippas, A.W., Slim, J., Bank, B., Sui, X., Kossman, S.E., Shenkenberg, T.D., Alexander, W.L., Price, K.A., Ley, J., Messina, D.N., Glasscock, J.I., Colevas, A.D., Cohen, E.E.W., Adkins, D., and Duncavage, E.J. (2024) Multicenter validation of an RNA-based assay to predict anti-PD-1 disease control in patients with recurrent or metastatic head and neck squamous cell carcinoma: the PREDAPT study. J Immunother Cancer. 12:e009573. DOI: 10.1136/jitc-2024-009573. PMID: 39489541; PMCID: PMC11535711.
Hiken, J., Earls, J., Flanagan, K.C., Wellinghoff, R.L., Ponder, M., Messina, D.N., Glasscock, J.I., and Duncavage, E.J. (2025) Analytical performance of OncoPrism-HNSCC, an RNA-based assay to inform immune checkpoint inhibitor treatment decisions for recurrent/metastatic head and neck squamous cell carcinoma. BMC Cancer. 25:21. DOI: 10.1186/s12885-024-13362-8. PMID: 39773366; PMCID: PMC11705923.
Marei, H.E., Hasan, A., Pozzoli, G., and Cenciarelli, C. (2023) Cancer immunotherapy with immune checkpoint inhibitors (ICIs): potential, mechanisms of resistance, and strategies for reinvigorating T cell responsiveness when resistance is acquired. Cancer Cell Int. 23:64. DOI:10.1186/s12935-023-02902-0. PMID: 37038154; PMCID: PMC10088229.
Moll, P., Ante, M., Seitz, A. and Reda, T. (2014) QuantSeq 3′ mRNA sequencing for RNA quantification. Nat Methods 11, i–iii (2014). DOI: 10.1038/nmeth.f.376
Written by Dr. Yvonne Göpel







