RNA sequencing (RNA-Seq) is a powerful tool that can be applied at various stages of the drug discovery and development workflow, from target identification to studying drug effects, mode-of-action, and monitoring disease progression and treatment responses. A thorough and careful experimental design is the most crucial aspect of an RNA-Seq experiment and key to ensuring meaningful results. Consulting specialists for experimental design and bioinformaticians for data analysis planning is essential for success and efficient use of resources. The following article explores key considerations for experimental design, data analysis, and follow-up studies.
Key considerations for designing RNA-Seq experiments for drug discovery and development
A clear understanding of your experimental requirements is essential for designing a successful RNA-Seq experiment in drug discovery studies. Careful planning can ensure your data effectively addresses your research questions, avoids costly pitfalls, and can potentially be mined as a general source of information for further projects. Considering several key questions for robust next generation sequencing (NGS) experiments upfront will help set you up for success (Fig. 1).
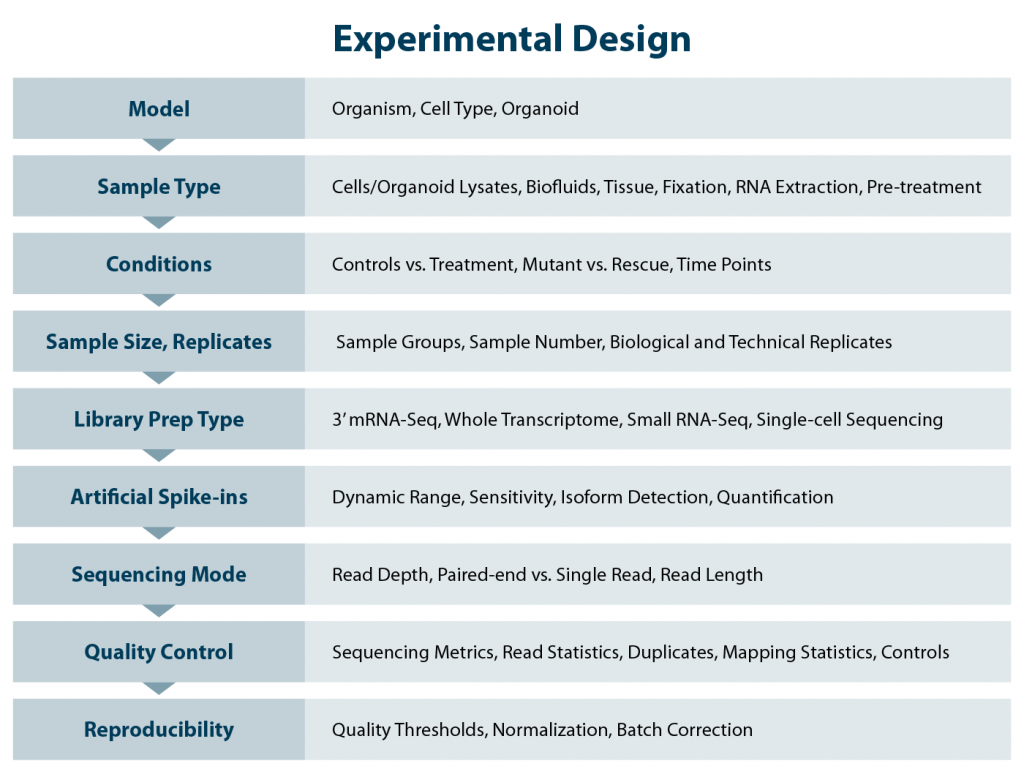
Aim of the study - hypothesis and objectives
Always start your study with a clear hypothesis and aim. Set the goals early to guide the experimental design from the chosen model system, the experimental conditions, plate layout, controls, library preparation method, sequencing setup and quality control parameters.
Based on your hypothesis and aim, you can decide on the best model system to address the research question.
- Are you interested in a specific target only, or can this project and and potential follow-up studies be used for data mining? Does the project benefit from a global, unbiased readout or is a targeted approach more suitable?
- What do you expect to find in terms of differential expression?
- Is the cell line or model system suitable to screen for the desired drug effects?
- Where do you expect variation, which level of variation, and how can you separate variability from genuine drug-induced effects?
- What type of data is needed to assess your hypothesis? Do you need quantitative data (e.g., gene expression) or qualitative data (e.g., coverage, isoforms, splice variation)?
These considerations have a direct influence on the wet lab workflow for the experiment, the data analysis and controls needed. Typical examples for RNA-Seq experiments during drug discovery studies are target identification, assessment of expression patterns in response to treatment, dose-response to compounds and drugs, drug combination effects, biomarker discovery and analysis, and mode-of-action studies.
Sample size and statistical power
The sample size for the drug discovery project has a significant impact on the quality and reliability of the results obtained in the study. Statistical power refers to the ability to identify genuine differential gene expression in naturally variable data sets. While there is an ideal sample size to ensure the optimal outcomes for statistic analysis, several factors influence the sample size, such as biological variation, complexity of the study, cost, and sample availability (Fig. 2). For example, it might be beneficial to include larger sample numbers or more replicates, however, for various sample types – especially precious patient samples from biobanks – this is virtually impossible. For other sample types, such as cell lines treated with various compounds, replicates and larger sample sizes can easily be managed.
- To ensure reliable results and robust conclusions, it is always beneficial to consult a bioinformatician or data expert to discuss the study and sample size limitations in regards to statistical power.
- Pilot studies are an excellent way to help determine sample size for the main experiment by assessing preliminary data on variability, and to test various conditions.
Replicates for drug discovery studies
The number of replicates is directly related to sample size and required to account for variability within and between experimental conditions.
- Biological Replicates are independent samples for the same experimental group / condition. They can account for natural variation between individuals, tissues, or cell populations. At least 3 biological replicates per condition are typically recommended. However, additional replicates should be considered when the samples can be easily sourced, e.g., cell lines, organoids, or to increase reliability when the variability in the experiment is high and dampens the signal. Ideally, between 4 – 8 replicates per sample group cover most experimental requirements.
- Technical Replicates can be included to assess technical variation, although biological replicates are more critical.
Replicates should also be considered in context of the desired data analysis (Table 1). Several bioinformatics tools require a minimum number of replicates for reliable data output (Schurch et al., 2016). Input from the bioinformaticians and data experts is highly valuable to optimize study design.
Table 1 | Biological vs. Technical Replicates.
| Biological Replicates | Technical Replicates | |
| Replicate | Different biological samples or entities (e.g., individuals, animals, cells) | The same biological sample, measured multiple times |
| Purpose | To assess biological variability and ensure findings are reliable and generalizable | To assess and minimize technical variation (variability of sequencing runs, lab workflows, environment) |
| Example | 3 different animals or cell samples in each experimental group (treatment vs. control) | 3 separate RNA sequencing experiments for the same RNA sample |
| Variability | Biological differences between individuals / subjects | Variation in measurement or workflow and environmental variability |
Experimental conditions and setup
A coherent experimental setup is the basis of a successful experiment. The number and type of the conditions assessed matter as much as sample collection, time points, concentrations, and the plate layout. Screening projects often use 384-well plate formats, while several steps for ‘omics’ workflows, e.g., RNA extractions, routinely use 96-well formats. Plate transfers should be planned in a way to ensure sample and experimental variability can be captured and batch effects can be corrected if needed.
- Cell type / model system
Choosing the appropriate cell type or (animal) model is paramount for your experiment. For cells or tissues, ensure that they are suitable to assess the drug’s effects in humans. Consider if additional cell lines from a different tissue are beneficial. In a later stage, is the animal model suitable or can organoids be used to assess complex systems? If you are working with patient samples, e.g., for retrospective studies, sample availability is a typical challenge.
- Conditions and treatment vs. control
Many RNA-Seq experiments during the drug discovery and development workflow compare samples from cells or patients treated with compounds, oligonucleotide therapeutics, or drug combinations vs. untreated control groups. When performing experiments in early discovery, cell lines are often an easily accessible model system of choice. Cell density, growth conditions, media, viability upon treatment, plate layout, and harvesting can affect variability within the experiment and should be tested upfront. Consider which are the best “no treatment” and “mock” controls for your experiment before entering larger experimental setups. Other common sample types such as blood and FFPE samples bring their own unique challenges. Explore more about transcriptomics from FFPE samples in our previous blog.
- Time points
Consider the time course of drug treatment and response. Drug effects on gene expression might vary over time, so multiple time points might be needed to catch the effect on the target. In addition, kinetic RNA sequencing with SLAMseq can be used to globally monitor RNA synthesis and decay rates. Choosing a kinetic RNA-Seq approach allows to distinguish primary from secondary drug effects and is particularly useful when candidates are assessed during mode-of-action studies. As multiple time points and replicates per sample group are needed to generate the relevant information, this approach is usually applied to select candidates only or for investigating specific drugs or combinations for more in-depth studies. This ensures that the total sample number remains manageable.
- Plate layout to enable batch correction
Batch effects refer to systematic, non-biological variations in the data that arise from how samples are collected and processed. Large-scale studies incur batch effects as samples cannot be collected and processed in parallel due to their number, time delays for treatments, and logistics challenges around the experiment. Samples are often grouped and processed in batches (so-called processing units) which can range from hundreds to thousands of samples at a time. Batch effects are expected for experiments that span across time, multiple sites, or large sample sets. Consider a design that allows to minimize batch effects and enables correction in silico. A variety of batch correction techniques and software tools are available that aim to remove batch effects and prevent confounding the downstream analysis. Learn more in BigOmic’s blog: What are Batch Effects in Omics Data and How to Correct Them.
- Experimental controls
Artificial spike-in controls, such as SIRVs, are valuable tools in RNA-Seq experiments that enable researchers to measure the performance of the complete assay, especially dynamic range, sensitivity, reproducibility, isoform detection, and quantification accuracy. Further, spike-in controls provide an internal standard that helps to quantify RNA levels between samples, normalize data, assess technical variability, and they can be used as quality control measure for large-scale experiments to ensure data consistency.
- Pilot studies
To mitigate risks and ensure reliable and relevant data for the main experiment, a pilot study with a representative sample subset is essential. This allows for validation of experimental parameters, wet lab and data analysis workflows. In addition, pilot studies enable a comparative evaluation of multiple methods before deciding on the best setup for the question at hand. Based on the pilot results, adjustments can be made before the start of the full-scale experiment.
Wet lab workflow: Library preparation for drug discovery
Aim and objectives of the study also drive the setup of the wet lab workflow or standard operating procedure throughout the discovery project. Several factors such as the type of data that is required, the RNA type of interest, the sample type and number influence the decision on the most effective NGS library preparation and pre-treatment regime.
For example, large-scale drug screens based on cultured cells with the aim to assess gene expression patterns or pathways, benefit from 3′-Seq approaches, with library preparation directly from lysates. This way, RNA extractions can be omitted, saving time and money and larger sample numbers can be handled efficiently ideally by early sample pooling. If isoforms, fusions, non-coding RNAs, or variants are of interest, whole transcriptome approaches are the method of choice, combined with mRNA enrichment or ribosomal rRNA depletion. In case whole blood samples or FFPE material are the input for the study, samples need to be extracted with care. Special emphasis is given to removing contaminants, abundant transcripts such as globin, genomic DNA, and to process low-quality and low-quantity samples ideally using streamlined and dedicated workflows.
- RNA extraction:
- Does your experiment require RNA extraction or can you use extraction-free RNA-Seq library preparation directly from the lysate, e.g., for studies using cell lines and large sample numbers?
- Does your RNA extraction kit recover the RNA species of interest, e.g., are small RNAs and microRNAs retained for small RNA biomarker discovery studies?
- Is the protocol suitable for your sample type, e.g., blood, biofluids, FFPE, and for potentially degraded RNA?
- Sample pre-treatment: Which type of pre-treatment do you need?
- gDNA removal,
- mRNA selection,
- rRNA depletion,
- removal of abundant transcripts,
- none: in-prep selection or enrichment panels post-library prep.
- Selecting the optimal library preparation protocol: The most suitable library preparation method depends on the sample type itself, the type of data you are interested in, sample numbers, input amounts, cost, and data analysis requirements. 3′ mRNA-Seq methods, such as QuantSeq and LUTHOR are ideal for gene expression and pathway analysis. For larger sample numbers, pooled library preparation approaches are particularly cost and time efficient. The analysis of microRNA biomarkers requires specialized small RNA-Seq library preparation protocols. Isoforms, fusions, and transcript variants can be assessed with whole transcriptome RNA-Seq, such as CORALL in combination with mRNA enrichment. For non-coding RNA analysis, rRNA depletion protocols are used prior to whole transcriptome RNA-Seq. It is recommended to select protocols that ensure strand specificity for highest accuracy and resolution of sense and anti-sense transcription (Mills et al., 2013).
- Choose a library preparation method that suits your experimental goals.
- Consider factors like enrichment, depletion, strand specificity, and pooling.
- If your experiment requires accurate quantification, consider incorporation of Unique Molecular Identifiers (UMIs). For example, if you are working with low input samples, low quality, or want to accurately quantify gene expression, fusions, or rare transcripts, UMIs are extremely beneficial.
- Choose a library preparation method suitable for your input quantity, e.g., when working with low inputs (<10 ng) or ultra-low inputs (<1 ng), ensure to select highly sensitive library preps.
- Consider the use of spike-in controls prior to library preparation for later quality control and normalization.
Sequencing and Quality Control
- Sequencing Platform:
- Select a sequencing platform based on cost, throughput, and read length requirements.
- Illumina platforms are commonly used for RNA-Seq. Platforms from other vendors, such as Element Biosciences, Singular Genomics, Ultima Genomics, or MGI can also be used. Depending on the manufacturer, most libraries can be either directly compatible or may need index adapter exchange or conversion allowing a versatile use of sequencing platforms without restricting the choice of library prep. Consult your supplier and instrument manufacturer for details.
- Outsourcing can deliver results fast and circumvent larger upfront investments in equipment and the need for trained personnel. Need tips to select a suitable sequencing service provider? Check our blog for more information: The Ultimate Guide to NGS Service Providers: Your Path to Drug Discovery Breakthroughs.
- Sequencing Depth: Determine the sequencing depth (number of reads per sample) based on the sample number, complexity, and the sensitivity required. Typically, for bulk RNA-Seq, 20-30 M Reads / Sample are sufficient for expression profiling. 3’ mRNA-Seq methods cover only the 3′ end of mRNA transcripts and thus require lower read depth (~3 – 5 M Reads / Sample). For large-scale screenings using pooled 3′ mRNA-Seq approaches, lower read depth is required, e.g. 200 K – 1 M Reads / Sample are a typical read out. Table 2 provides an overview of the recommended sequencing depth for various applications.
- Determine the appropriate sequencing depth to achieve sufficient coverage for your analysis.
- Consult a bioinformatician or data analysis expert in case of doubt and perform a pilot experiment to check the sequencing parameters.
Table 2 | Sequencing Read Recommendations by Application.

- Read length and sequencing mode: The optimal read length and sequencing mode depend on the application and architecture of the library. Most applications can be sequenced in single read mode, e.g., SR100. When your libraries contain inline indices or UMIs, full or partial paired-end reads may be required. Ensure to select the appropriate sequencing mode to accommodate inline indices and UMI read-out.
- For gene expression / transcriptome profiling (whole transcriptome or 3′ mRNA-Seq) single read (SR) 75 – 100 are typically used. Paired-end sequencing is possible; however, for 3′-Seq, Read 2, which begins with the homopolymer stretch corresponding to the oligo(dT) primer binding to the poly(A) tail, should be discarded.
- Alternative splicing, rare transcripts, variants, and assemblies benefit from longer, paired-end reads covering splice sites and spanning larger regions. Typical read modes are PE75, PE100, and PE150.
- Small RNA analysis uses dedicated library types. These short libraries are typically sequenced in SR50 or SR75 mode which covers the entire sequence. In case UMIs are included in your small RNA libraries, ensure the read length is suitable to cover the small RNA insert and the UMI sequence.
- Quality Control: It is recommended to use appropriate in-sequencing controls and to check the quality of the sequencing run before proceeding to data QC and analysis. To learn more about Quality Control and Primary Data Analysis visit our Lexicon blog.
Data Analysis
In drug discovery RNA-Seq experiments, the primary objective is to produce high-quality data that accurately reflects drug-induced gene expression changes, aiding in biomarker and target identification. To avoid investing resources in analyzing compromised data, rigorous quality assessment is crucial. Early detection of issues ensures sound biological conclusions, as flawed data inevitably leads to unreliable results during subsequent analyses.
- Always quality control read statistics, e.g., with FASTQC (Andrews, 2010).
- Incorporating artificial spike-ins into RNA-Seq experiments at a low read percentage provides further quality insights. Spike-ins enable robust data comparison across time points and locations, and facilitate rapid initial quality control. Artificial Spike-in controls serve as a reliable proxy for evaluating library generation and sequencing workflow performance. Unexpected results in these controls can pinpoint issues related to sample quality, cross-contamination, or workflow discrepancies, allowing for timely adjustments of the operating procedure.
- Once QC checks are passed, the data can be processed as required. Primary and secondary analysis cover read processing, alignment and quantification, and provide the input needed for biological insights. Consider which normalization and corrections need to be applied to the data sets to ensure consistency across the complete experiment.
- Data analysis aims to convert sequencing data into knowledge within the respective biological context. Both, statistical know-how and expertise in (cancer) biology are required to interpret the results. Various analyses are possible depending on the experimental goal: most common applications aim to identify expression patterns, pathways and biomarkers indicative of a drugs action, disease progression or treatment response.
- If available, integrate RNA-Seq data with other omics data (proteomics, metabolomics) for a more comprehensive understanding of drug effects. Sophisticated tools and platforms are available for integration of omics data and data interpretation, e.g., the Omics Playground from BigOmics Analytics. These tools can help identify patterns, networks, draw conclusions, suggest modes-of-action for validation, and synergies for follow-up studies.
Experimental Validation
Part of the experimental validation includes assessing whether the observed changes in gene expression are consistent with known drug mechanisms, as well as confirming the data with independent methods, e.g., qPCR, cell-based assays, and follow-up experiments elucidating the underlying molecular regulation of the drug effect. It is essential to confirm dose-dependency, to identify primary and secondary effects, and assess potential resistance mechanisms. Validation studies should also undergo a rigoros planning phase to ensure suitability of the approach and unbiased assessment of the results to confirm previous observations. Sample randomization or blind testing can be used to minimize expectation bias.
Summary
RNA-Seq is a powerful tool for drug discovery with highly versatile applications. This comprehensive guide to designing and executing successful RNA-Seq experiments for drug discovery and development emphasizes the importance of meticulous planning.
- Clearly defined hypotheses and objectives guide the entire experimental process.
- Optimal sample size and statistical power ensure robust statistical analysis and can be determined in consultation with bioinformaticians and by conducting pilot studies.
- Using biological replicates to account for data variability and technical replicates when necessary is key for reliable and consistent results.
- Careful selection of cell types, treatments, time points, and plate layouts ensures the experimental conditions are appropriate to assess the hypothesis.
- The choice of sample processing workflows and library preparation protocols is based on sample type and research goals, including the use of spike-in controls.
- Selecting appropriate sequencing platforms, read depths, and modes, and implementing rigorous quality checks ensures optimized sequencing setup.
- Employing appropriate bioinformatics tools for QC, normalization, analysis and the integration with other omics data are key to biologically sound conclusions and meaningful results.
- Experimental validation by confirming RNA-Seq results with independent methods like qPCR and cell-based assays adds confidence to the study and validates results from NGS experiments.
References
Andrews S. (2010). FastQC: a quality control tool for high throughput sequence data. Available online at: http://www.bioinformatics.babraham.ac.uk/projects/fastqc
Campbell, J.D., Liu G., Luo, L., Xiao, J., Gerrein, J., Juan-Guardela, B., Tedrow, J., Alekseyev, Y.O., Yang, I.V., Correll, M., Geraci, M., Quackenbush, J., Sciurba, F., Schwartz, D.A., Kaminski, N., Johnson, W.E., Monti, S., Spira, A., Beane, J., Lenburg, M.E. (2015) Assessment of microRNA differential expression and detection in multiplexed small RNA sequencing data. RNA. 21(2):164-71. DOI: 10.1261/rna.046060.114. PMID: 25519487; PMCID: PMC4338344
Liu, Y., Zhou, J., and Kevin P. White, K. P. (2014) RNA-seq differential expression studies: more sequence or more replication? Bioinformatics, 30(3): 301–304, DOI: 10.1093/bioinformatics/btt688
Mills J.D., Kawahara Y., Janitz M. (2013) Strand-Specific RNA-Seq Provides Greater Resolution of Transcriptome Profiling. Curr Genomics. 14(3):173-81. DOI: 10.2174/1389202911314030003. PMID: 24179440; PMCID: PMC3664467
Moll, P., Ante, M., Seitz, A. and Reda, T. (2014) QuantSeq 3′ mRNA sequencing for RNA quantification. Nat Methods 11, i–iii (2014). DOI: 10.1038/nmeth.f.376
Schurch, N..J, Schofield, P., Gierliński, M., Cole, C., Sherstnev, A., Singh, V., Wrobel, N., Gharbi, K., Simpson, G.G., Owen-Hughes, T., Blaxter, M., Barton, G.J. (2016) How many biological replicates are needed in an RNA-seq experiment and which differential expression tool should you use? RNA 22(6):839-51. DOI: 10.1261/rna.053959.115.
Written by Dr. Yvonne Göpel









