Let us help you to improve your SARS-CoV-2 sequencing workflows
- Reliable method to detect and trace viral variants
- Fast and easy ready-to-sequence library prep for Illumina
CORALL and SARS-CoV-2 Library Enrichment
The hybridization / capture enrichment workflow enables whole transcriptome sequencing of SARS-CoV-2 variants by post-library prep enrichment of viral sequences from CORALL RNA-Seq libraries. The complete workflow enriches viral reads up to 99 % for NGS approaches and robustly captures even variants with significant mutations and rearrangements.
Performance
- Epidemiologic surveillance sequencing for variant identification and tracking.
- Monitoring of viral evolution, spread, and transmission routes.
Featured Products
- CORALL RNA-Seq Library Preparation Kit (Cat. No. 095)
- myBaits Expert SARS-CoV-2 (Daciel Arbor Biosciences, Cat. No. 308716.v5)
- Re-Amplification Kit (Cat. No. 080)
Overview
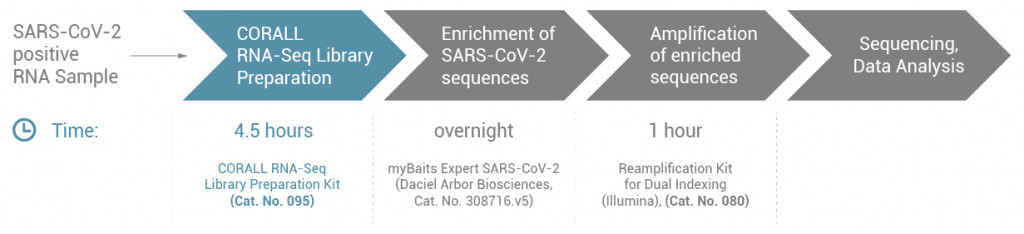
In 4.5 hours, CORALL Total RNA-Seq libraries were prepared from a SARS-CoV-2 positive RNA sample. The myBaits Expert SARS-CoV-2 Kit (Daciel Arbor Biosciences) was used for enrichment of SARS-CoV-2 sequences, followed by amplification of the enriched libraries (Fig. 1).

Performance
Enrichment of viral sequences from CORALL RNA-Seq libraries with myBaits Expert SARS-CoV-2 increases the fraction of SARS-CoV-2 mapped reads from ~0.1 % up to ~99 % and shows even coverage distribution at a depth of 1 Mb (Fig. 2).
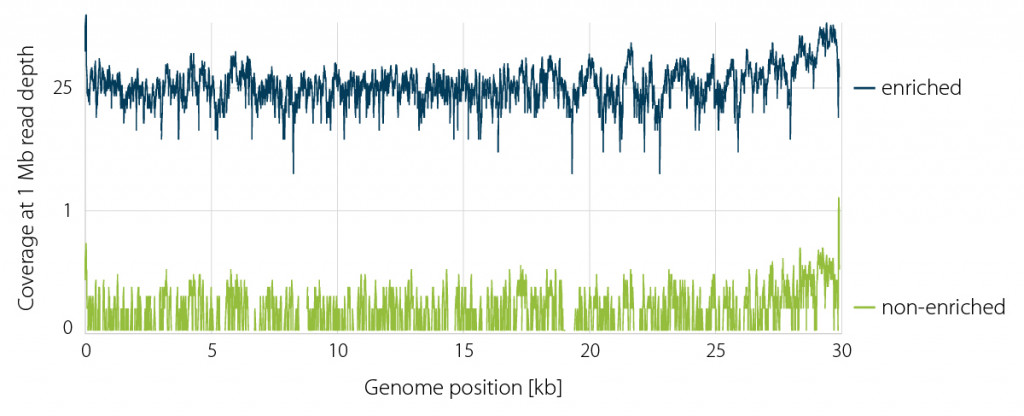
Figure 2 | Normalized CORALL coverage of the complete SARS-CoV-2 genome prior (green) and post enrichment (blue) with a hybridization / capture enrichment panel (myBaits Expert SARS-CoV-2, Cat. No. 308716.v5, Daciel Arbor Biosciences).
Figure 2 shows the per base SARS-CoV-2 genome coverage without enrichment at 0.02±0.03 x using 1 Mio reads. The majority of reads belonged to bacteria (ca. 95 %) and the human host (ca. 5 %). Following enrichment, SARS-CoV-2 genome coverage increased to 33±24 x with 95.7% of bases exceeding a coverage 10x which is sufficient for confident mutation calling.
Buy from the Webstore
Contact us for product information or an introductory offer
For more information please contact us at support@lexogen.com




