Step 1: Reverse Transcription

![]()
The kit uses total RNA as input, hence no prior poly(A) enrichment or rRNA depletion is needed.
![]()
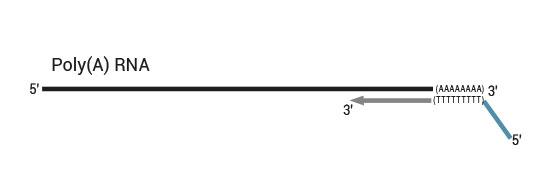
Library generation starts with oligodT priming containing the Ion Torrent-specific P1 adapter in the 5`part of the oligodT primer.
Step 2: Removal of RNA
![]()
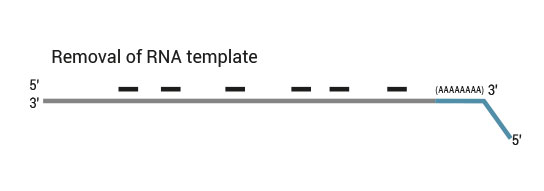
After first strand synthesis the RNA is removed.
Step 3: Second Strand Synthesis
![]()
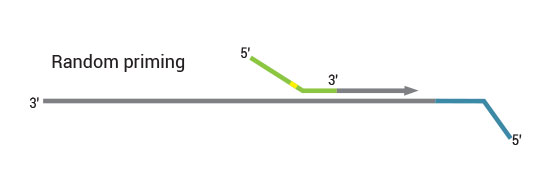
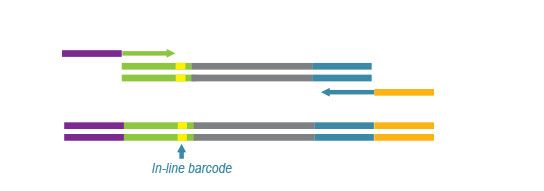
Second strand synthesis is initiated by random priming and a DNA polymerase. The random primer contains the Ion Torrent-specific A adapter in the 5’ end of the random primer and up to 48 in-line barcodes can be introduced.
![]()
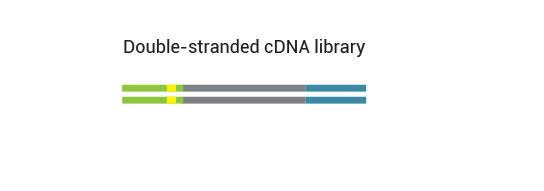
No purification is required between first and second strand synthesis. Second strand synthesis is followed by a magnetic bead-based purification step rendering the protocol compatible with automation.
Step 4: Library Amplification
![]()
During the library amplification step the sequences required for colony formation are introduced.



