Scalable and Rapid Blood Transcriptomics with QuantSeq-Pool for Blood
High-throughput Expression Profiling for Blood
Lexogen’s QuantSeq-Pool for Blood enables massively multiplexed blood RNA-Seq for fast and cost-effective gene expression screening of whole blood samples. Elevate blood transcriptomics with robust and reliable pooled gene expression data, unparalleled scalability, and significant cost and plasticware savings.
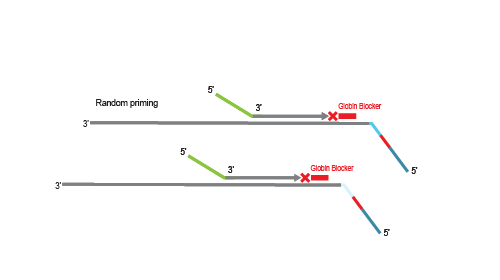
QuantSeq-Pool uses sample barcoding and early pooling to maximize sample throughput up to 36,864 samples in one sequencing run. The established 3’ mRNA-Seq library generation uses QuantSeq technology and promotes high-throughput screenings for straightforward mRNA-focused readouts. Integrated globin blockers efficiently prevent the generation of library fragments from globin mRNAs by blocking their extension during second strand synthesis.
QuantSeq-Pool for Blood is available upon request!
Send us your request to access Lexogen’s solution for High-throughput Expression Profiling for Blood!
Workflow
The kit uses total RNA as input, hence no prior poly(A) enrichment or rRNA depletion is needed. For balanced read distribution in the sequencing experiment the RNA input
amount for reverse transcription needs to be adjusted and normalized across all
samples that shall be pooled.
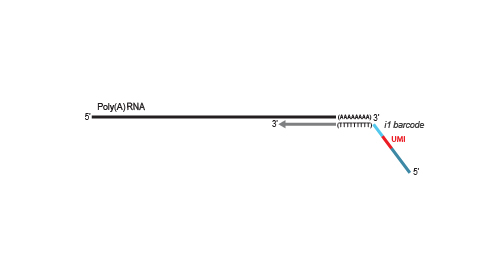
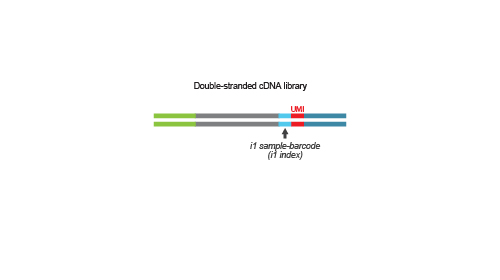
Library generation is initiated by oligo(dT) priming. The primer already contains a partial Illumina-specific Read 2 linker sequence, Unique Molecular Identifiers (UMIs), and an i1 sample-barcode to uniquely label each sample.
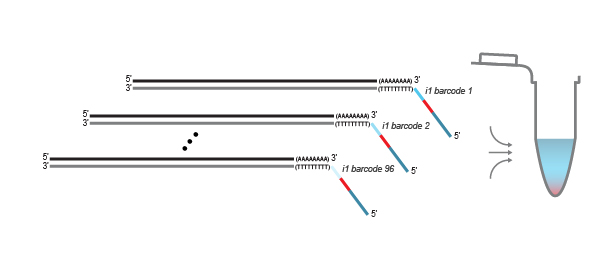
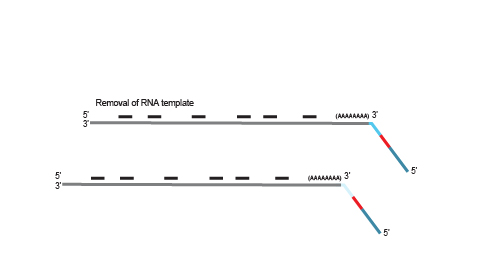
After first strand synthesis, individual samples are combined by pooling. The pool is purified to decrease the volume for subsequent steps. All further reactions are carried out in batch
on the combined samples to save time and effort.
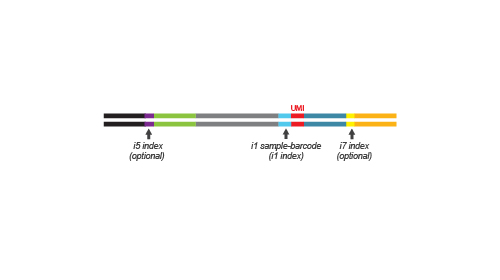
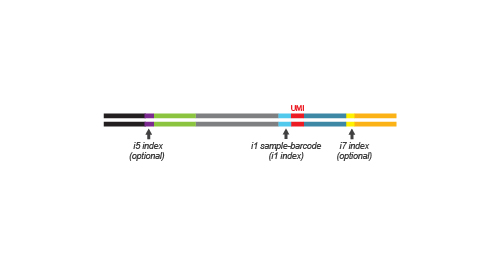
During the library amplification step sequences required for cluster generation are introduced. Optionally, i5 and i7 indices can be introduced. These additional indices are required for multiplexing more than 96 samples.
Multiplexing capacities can reach up to 9,216 samples when 96 additional i5 / i7 Unique Dual Indices (UDIs) are used or up to 36,864 samples when 384 UDIs are used.
The final purification elutes ready-to-sequence library pools of up to 96 samples which directly correspond to the final lane pool for a 96-plex sequencing experiment.
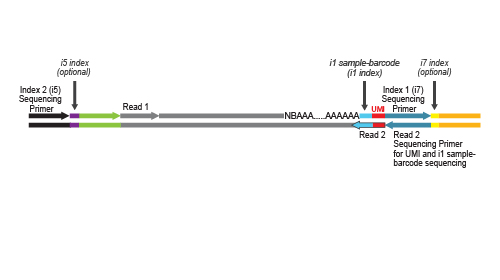
NGS reads are generated towards the poly(A) tail and directly correspond to the mRNA sequence. The length distribution of QuantSeq-
Pool libraries supports all sequencing lengths. Read 2 is required to read out the sequence information for the UMI and the i1 sample-barcode. Therefore, at least a limited Read 2 with a read length between 18 and 22 nucleotides is mandatory for sequencing QuantSeq-Pool libraries.
First time user of QuantSeq-Pool for Blood?
First Time User? We’re excited to offer you an exclusive introductory offer.